Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1745 [new locus tag: SACOL_RS08930 ]
- pan locus tag?: SAUPAN004324000
- symbol: pyk
- pan gene symbol?: pykA
- synonym:
- product: pyruvate kinase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1745 [new locus tag: SACOL_RS08930 ]
- symbol: pyk
- product: pyruvate kinase
- replicon: chromosome
- strand: -
- coordinates: 1781926..1783683
- length: 1758
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238347 NCBI
- RefSeq: YP_186581 NCBI
- BioCyc: see SACOL_RS08930
- MicrobesOnline: 913191 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741ATGAGAAAAACTAAAATTGTATGTACAATTGGACCAGCTTCAGAATCAGAAGAAATGATT
GAGAAATTAATCAATGCTGGTATGAACGTTGCACGATTAAACTTTTCACATGGTAGTCAT
GAAGAGCATAAAGGTAGAATTGATACAATTCGTAAAGTAGCTAAAAGATTAGACAAAATT
GTAGCAATTTTATTAGATACAAAAGGTCCAGAAATTCGTACGCATAATATGAAAGACGGT
ATCATTGAACTTGAACGTGGCAACGAAGTTATTGTTAGCATGAATGAAGTTGAAGGAACA
CCTGAAAAGTTCTCAGTAACATATGAAAACTTAATTAACGATGTTCAAGTAGGTTCATAC
ATTTTACTTGATGATGGCTTAATTGAATTACAAGTTAAAGATATTGACCATGCTAAAAAA
GAAGTTAAATGTGATATTTTAAACTCTGGTGAGCTTAAAAACAAAAAAGGTGTTAACTTA
CCTGGCGTAAGAGTAAGTTTACCTGGTATTACAGAAAAAGATGCTGAAGATATCCGTTTC
GGTATTAAAGAAAATGTTGACTTCATTGCAGCAAGTTTCGTACGTCGTCCTAGTGATGTT
TTAGAAATTCGTGAAATTTTAGAAGAACAAAAAGCTAACATTTCAGTATTCCCTAAAATT
GAAAACCAAGAAGGTATTGATAATATTGCGGAAATTCTTGAAGTGTCTGATGGTTTAATG
GTTGCACGTGGTGACATGGGTGTTGAAATTCCACCTGAAAAAGTACCAATGGTTCAAAAA
GATTTAATCAGACAATGTAACAAATTAGGTAAACCAGTTATTACAGCTACACAAATGTTA
GATTCTATGCAACGTAACCCACGTGCTACACGTGCAGAAGCTAGTGACGTTGCCAACGCA
ATCTATGATGGTACAGATGCAGTAATGTTATCTGGTGAAACTGCTGCTGGTTTATATCCT
GAAGAAGCTGTTAAAACAATGAGAAATATTGCTGTATCAGCTGAAGCAGCCCAAGATTAC
AAAAAGTTATTGTCAGATCGTACTAAATTAGTTGAAACTTCATTAGTGAATGCTATCGGT
ATTTCGGTTGCACATACAGCTTTAAACTTAAATGTTAAAGCAATTGTAGCTGCTACTGAA
AGTGGTTCAACGGCACGTACTATCTCCAAATATCGTCCACATTCAGACATTATTGCGGTG
ACTCCAAGTGAAGAAACTGCACGTCAATGTTCAATTGTTTGGGGAGTTCAACCTGTAGTT
AAAAAAGGACGTAAGAGTACAGATGCATTGTTAAACAATGCAGTTGCAACAGCTGTTGAA
ACTGGTAGAGTATCTAATGGTGATTTAATCATTATTACTGCTGGTGTACCAACTGGTGAA
ACTGGAACTACTAATATGATGAAAATCCACCTAGTTGGTGACGAAATTGCTAATGGTCAA
GGTATTGGACGTGGATCAGTTGTTGGTACTACGTTAGTTGCTGAAACTGTTAAAGATTTA
GAAGGTAAAGATTTATCTGACAAAGTTATCGTTACTAACTCAATCGATGAAACGTTTGTA
CCTTATGTAGAAAAAGCTTTAGGCTTAATTACAGAAGAAAATGGTATTACATCACCAAGT
GCAATTGTTGGTTTAGAAAAAGGTATTCCAACAGTTGTAGGTGTAGAAAAAGCTGTTAAA
AACATAAGCAATAACATGTTAGTTACGATTGATGCTGCTCAAGGTAAAATCTTTGAAGGA
TATGCAAACGTACTATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1758
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1745 [new locus tag: SACOL_RS08930 ]
- symbol: Pyk
- description: pyruvate kinase
- length: 585
- theoretical pI: 4.99555
- theoretical MW: 63101.9
- GRAVY: -0.127863
⊟Function[edit | edit source]
- reaction: EC 2.7.1.40? ExPASyPyruvate kinase ATP + pyruvate = ADP + phosphoenolpyruvate
- TIGRFAM: Energy metabolism Glycolysis/gluconeogenesis pyruvate kinase (TIGR01064; EC 2.7.1.40; HMM-score: 619.1)and 4 moreEnergy metabolism Glycolysis/gluconeogenesis phosphoenolpyruvate synthase (TIGR01418; EC 2.7.9.2; HMM-score: 56.3)phosphoenolpyruvate-protein phosphotransferase (TIGR01417; EC 2.7.3.9; HMM-score: 27.2)2-dehydro-3-deoxyglucarate aldolase (TIGR03239; EC 4.1.2.20; HMM-score: 15.6)2,4-dihydroxyhept-2-ene-1,7-dioic acid aldolase (TIGR02311; EC 4.1.2.-; HMM-score: 14.4)
- TheSEED :
- Phosphohistidine swiveling domain
- Pyruvate kinase (EC 2.7.1.40)
Carbohydrates Central carbohydrate metabolism Glycolysis and Gluconeogenesis Pyruvate kinase (EC 2.7.1.40)and 3 more - PFAM: TIM_barrel (CL0036) PK; Pyruvate kinase, barrel domain (PF00224; HMM-score: 492.9)and 6 moreno clan defined PK_C; Pyruvate kinase, alpha/beta domain (PF02887; HMM-score: 137.2)Leu-IlvD (CL0364) PEP-utilizers; PEP-utilising enzyme, mobile domain (PF00391; HMM-score: 61.7)HTH (CL0123) IF2_N; Translation initiation factor IF-2, N-terminal region (PF04760; HMM-score: 18.4)TIM_barrel (CL0036) HpcH_HpaI; HpcH/HpaI aldolase/citrate lyase (PF03328; HMM-score: 14.2)Cellulase; Cellulase (glycosyl hydrolase family 5) (PF00150; HMM-score: 13)Mur_ligase_C_sf (CL0794) Mur_ligase_C; Mur ligase, glutamate ligase domain (PF02875; HMM-score: 11.9)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: K+, Mg2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9063
- Cytoplasmic Membrane Score: 0.0119
- Cell wall & surface Score: 0.0015
- Extracellular Score: 0.0803
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.012748
- TAT(Tat/SPI): 0.000555
- LIPO(Sec/SPII): 0.001816
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MRKTKIVCTIGPASESEEMIEKLINAGMNVARLNFSHGSHEEHKGRIDTIRKVAKRLDKIVAILLDTKGPEIRTHNMKDGIIELERGNEVIVSMNEVEGTPEKFSVTYENLINDVQVGSYILLDDGLIELQVKDIDHAKKEVKCDILNSGELKNKKGVNLPGVRVSLPGITEKDAEDIRFGIKENVDFIAASFVRRPSDVLEIREILEEQKANISVFPKIENQEGIDNIAEILEVSDGLMVARGDMGVEIPPEKVPMVQKDLIRQCNKLGKPVITATQMLDSMQRNPRATRAEASDVANAIYDGTDAVMLSGETAAGLYPEEAVKTMRNIAVSAEAAQDYKKLLSDRTKLVETSLVNAIGISVAHTALNLNVKAIVAATESGSTARTISKYRPHSDIIAVTPSEETARQCSIVWGVQPVVKKGRKSTDALLNNAVATAVETGRVSNGDLIIITAGVPTGETGTTNMMKIHLVGDEIANGQGIGRGSVVGTTLVAETVKDLEGKDLSDKVIVTNSIDETFVPYVEKALGLITEENGITSPSAIVGLEKGIPTVVGVEKAVKNISNNMLVTIDAAQGKIFEGYANVL
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3] [4] [5]
- quantitative data / protein copy number per cell: 4298 [6]
- interaction partners:
SACOL1760 (ackA) acetate kinase [7] (data from MRSA252) SACOL1385 (acnA) aconitate hydratase [7] (data from MRSA252) SACOL0660 (adhP) alcohol dehydrogenase [7] (data from MRSA252) SACOL0451 (ahpF) alkyl hydroperoxide reductase subunit F [7] (data from MRSA252) SACOL1478 (ald1) alanine dehydrogenase [7] (data from MRSA252) SACOL2657 (arcA) arginine deiminase [7] (data from MRSA252) SACOL2656 (arcB2) ornithine carbamoyltransferase [7] (data from MRSA252) SACOL0557 (cysK) cysteine synthase [7] (data from MRSA252) SACOL1637 (dnaK) molecular chaperone DnaK [7] (data from MRSA252) SACOL1587 (efp) elongation factor P [7] (data from MRSA252) SACOL0842 (eno) phosphopyruvate hydratase [7] (data from MRSA252) SACOL0634 (eutD) phosphotransacetylase [7] (data from MRSA252) SACOL2622 (fdaB) fructose-1,6-bisphosphate aldolase [7] (data from MRSA252) SACOL1329 (femC) glutamine synthetase [7] (data from MRSA252) SACOL1199 (ftsZ) cell division protein FtsZ [7] (data from MRSA252) SACOL0593 (fusA) elongation factor G [7] (data from MRSA252) SACOL0838 (gapA1) glyceraldehyde 3-phosphate dehydrogenase [7] (data from MRSA252) SACOL0877 (gcvH) glycine cleavage system protein H [7] (data from MRSA252) SACOL2145 (glmS) glucosamine--fructose-6-phosphate aminotransferase [7] (data from MRSA252) SACOL1554 (gnd) 6-phosphogluconate dehydrogenase [7] (data from MRSA252) SACOL0461 (guaA) GMP synthase [7] (data from MRSA252) SACOL0460 (guaB) inosine-5'-monophosphate dehydrogenase [7] (data from MRSA252) SACOL1741 (icd) isocitrate dehydrogenase [7] (data from MRSA252) SACOL1477 (ilvA1) threonine dehydratase [7] (data from MRSA252) SACOL2618 (ldh2) L-lactate dehydrogenase [7] (data from MRSA252) SACOL2623 (mqo2) malate:quinone oxidoreductase [7] (data from MRSA252) SACOL0792 (nrdE) ribonucleotide-diphosphate reductase subunit alpha [7] (data from MRSA252) SACOL1102 (pdhA) pyruvate dehydrogenase complex E1 component subunit alpha [7] (data from MRSA252) SACOL1104 (pdhC) branched-chain alpha-keto acid dehydrogenase E2 [7] (data from MRSA252) SACOL1105 (pdhD) dihydrolipoamide dehydrogenase [7] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [7] (data from MRSA252) SACOL0204 (pflB) formate acetyltransferase [7] (data from MRSA252) SACOL0966 (pgi) glucose-6-phosphate isomerase [7] (data from MRSA252) SACOL0839 (pgk) phosphoglycerate kinase [7] (data from MRSA252) SACOL1091 (ptsH) phosphocarrier protein HPr [7] (data from MRSA252) SACOL0584 (rplA) 50S ribosomal protein L1 [7] (data from MRSA252) SACOL2236 (rplB) 50S ribosomal protein L2 [7] (data from MRSA252) SACOL2227 (rplE) 50S ribosomal protein L5 [7] (data from MRSA252) SACOL2224 (rplF) 50S ribosomal protein L6 [7] (data from MRSA252) SACOL0585 (rplJ) 50S ribosomal protein L10 [7] (data from MRSA252) SACOL0583 (rplK) 50S ribosomal protein L11 [7] (data from MRSA252) SACOL0586 (rplL) 50S ribosomal protein L7/L12 [7] (data from MRSA252) SACOL2220 (rplO) 50S ribosomal protein L15 [7] (data from MRSA252) SACOL2232 (rplP) 50S ribosomal protein L16 [7] (data from MRSA252) SACOL1257 (rplS) 50S ribosomal protein L19 [7] (data from MRSA252) SACOL1702 (rplU) 50S ribosomal protein L21 [7] (data from MRSA252) SACOL2234 (rplV) 50S ribosomal protein L22 [7] (data from MRSA252) SACOL2237 (rplW) 50S ribosomal protein L23 [7] (data from MRSA252) SACOL0545 (rplY) 50S ribosomal protein L25/general stress protein Ctc [7] (data from MRSA252) SACOL2213 (rpoA) DNA-directed RNA polymerase subunit alpha [7] (data from MRSA252) SACOL0588 (rpoB) DNA-directed RNA polymerase subunit beta [7] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [7] (data from MRSA252) SACOL2233 (rpsC) 30S ribosomal protein S3 [7] (data from MRSA252) SACOL1769 (rpsD) 30S ribosomal protein S4 [7] (data from MRSA252) SACOL2222 (rpsE) 30S ribosomal protein S5 [7] (data from MRSA252) SACOL0437 (rpsF) 30S ribosomal protein S6 [7] (data from MRSA252) SACOL2206 (rpsI) 30S ribosomal protein S9 [7] (data from MRSA252) SACOL2214 (rpsK) 30S ribosomal protein S11 [7] (data from MRSA252) SACOL2215 (rpsM) 30S ribosomal protein S13 [7] (data from MRSA252) SACOL2230 (rpsQ) 30S ribosomal protein S17 [7] (data from MRSA252) SACOL0009 (serS) seryl-tRNA synthetase [7] (data from MRSA252) SACOL0095 (spa) immunoglobulin G binding protein A precursor [7] (data from MRSA252) SACOL1262 (sucC) succinyl-CoA synthetase subunit beta [7] (data from MRSA252) SACOL1722 (tig) trigger factor [7] (data from MRSA252) SACOL0840 (tpiA) triosephosphate isomerase [7] (data from MRSA252) SACOL1155 (trxA) thioredoxin [7] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [7] (data from MRSA252) SACOL0521 hypothetical protein [7] (data from MRSA252) SACOL0564 pyridoxal biosynthesis lyase PdxS [7] (data from MRSA252) SACOL0815 ribosomal subunit interface protein [7] (data from MRSA252) SACOL1020 hypothetical protein [7] (data from MRSA252) SACOL1437 CSD family cold shock protein [7] (data from MRSA252) SACOL1759 universal stress protein [7] (data from MRSA252) SACOL1952 ferritins family protein [7] (data from MRSA252) SACOL2173 alkaline shock protein 23 [7] (data from MRSA252) SACOL2569 1-pyrroline-5-carboxylate dehydrogenase [7] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: CcpA regulon
CcpA (TF) important in Carbon catabolism; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
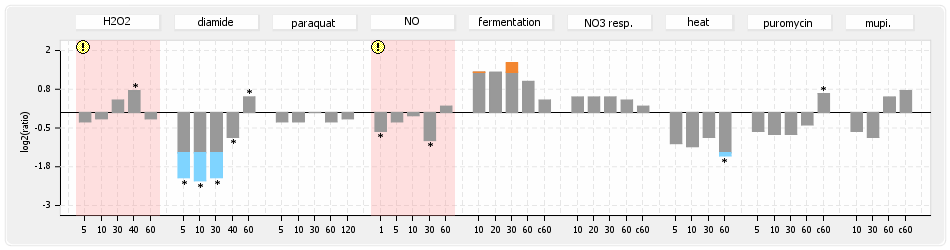
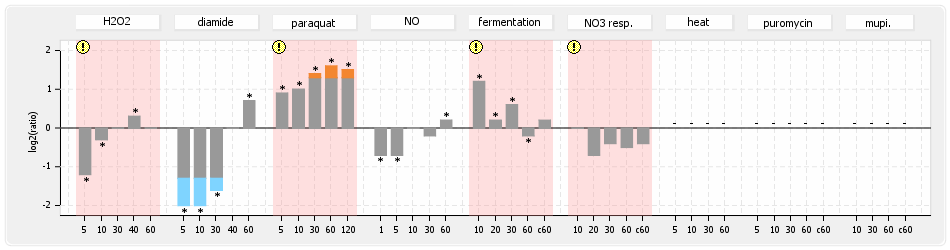
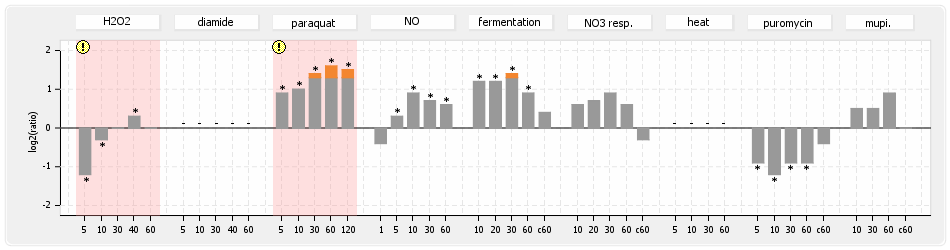
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 58.55 h [8]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Annette Dreisbach, Kristina Hempel, Girbe Buist, Michael Hecker, Dörte Becher, Jan Maarten van Dijl
Profiling the surfacome of Staphylococcus aureus.
Proteomics: 2010, 10(17);3082-96
[PubMed:20662103] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 7.00 7.01 7.02 7.03 7.04 7.05 7.06 7.07 7.08 7.09 7.10 7.11 7.12 7.13 7.14 7.15 7.16 7.17 7.18 7.19 7.20 7.21 7.22 7.23 7.24 7.25 7.26 7.27 7.28 7.29 7.30 7.31 7.32 7.33 7.34 7.35 7.36 7.37 7.38 7.39 7.40 7.41 7.42 7.43 7.44 7.45 7.46 7.47 7.48 7.49 7.50 7.51 7.52 7.53 7.54 7.55 7.56 7.57 7.58 7.59 7.60 7.61 7.62 7.63 7.64 7.65 7.66 7.67 7.68 7.69 7.70 7.71 7.72 7.73 7.74 7.75 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)