Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1390 [new locus tag: SACOL_RS07075 ]
- pan locus tag?: SAUPAN003742000
- symbol: parC
- pan gene symbol?: parC
- synonym:
- product: DNA topoisomerase IV subunit A
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1390 [new locus tag: SACOL_RS07075 ]
- symbol: parC
- product: DNA topoisomerase IV subunit A
- replicon: chromosome
- strand: +
- coordinates: 1397970..1400372
- length: 2403
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3237159 NCBI
- RefSeq: YP_186243 NCBI
- BioCyc: see SACOL_RS07075
- MicrobesOnline: 912849 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861
1921
1981
2041
2101
2161
2221
2281
2341
2401GTGAGTGAAATAATTCAAGATTTATCACTTGAAGATGTTTTAGGTGATCGCTTTGGAAGA
TATAGTAAATATATTATTCAAGAGCGTGCATTGCCAGATGTTCGTGATGGTTTAAAACCA
GTACAACGTCGTATTTTATATGCAATGTATTCAAGTGGTAATACACACGATAAAAATTTC
CGTAAAAGTGCGAAAACAGTCGGTGATGTTATTGGTCAATATCATCCACATGGAGACTCC
TCAGTGTACGAAGCAATGGTCCGTTTAAGTCAAGACTGGAAGTTACGACATGTCTTAATA
GAAATGCATGGTAATAATGGTAGTATCGATAATGATCCGCCAGCGGCAATGCGTTACACT
GAAGCTAAGTTAAGCTTACTAGCTGAAGAGTTATTACGTGATATTAATAAAGAGACAGTT
TCTTTCATTCCAAACTATGATGATACGACACTCGAACCAATGGTATTGCCATCAAGATTT
CCTAACTTACTAGTGAATGGTTCTACAGGTATATCTGCAGGTTACGCGACAGATATACCA
CCACATAATTTAGCTGAAGTGATTCAAGCAACACTTAAATATATTGATAATCCGGATATT
ACAGTCAATCAATTAATGAAATATATTAAAGGTCCTGATTTTCCAACTGGTGGTATTATT
CAAGGTATTGATGGTATTAAAAAAGCTTATGAATCAGGTAAAGGTAGAATTATAGTTCGT
TCTAAAGTTGAAGAAGAAACTTTACGCAATGGACGTAAACAGTTAATTATTACTGAAATT
CCATATGAAGTGAACAAAAGTAGCTTAGTAAAACGTATCGATGAATTACGTGCTGACAAA
AAAGTCGATGGTATCGTTGAAGTACGTGATGAAACTGATAGAACTGGTTTACGAATAGCA
ATTGAATTGAAAAAAGATGTGAACAGTGAATCAATCAAAAATTATCTTTATAAAAACTCT
GATTTACAGATTTCATATAATTTCAACATGGTCGCTATTAGTGATGGTCGTCCAAAATTG
ATGGGTATTCGTCAAATTATAGATAGTTATTTGAATCACCAAATTGAGGTTGTTGCAAAT
AGAACGAAGTTTGAATTAGATAATGCAGAAAAACGTATGCATATCGTTGAAGGTTTGATT
AAAGCGTTGTCAATTTTAGATAAAGTAATCGAATTGATTCGTAGCTCTAAAAACAAGCGT
GACGCTAAAGAAAACCTTATCGAAGTATACGAGTTCACAGAAGAACAGGCTGAAGCAATT
GTAATGTTACAGTTATATCGTTTAACAAATACTGACATAGTTGCGCTTGAAGGTGAACAT
AAAGAACTTGAAGCATTAATCAAACAATTACGTCATATTCTTGATAACCATGATGCATTA
TTGAATGTCATAAAAGAAGAATTGAATGAAATTAAAAAGAAATTCAAATCTGAACGACTG
TCTTTAATTGAAGCAGAAATTGAAGAAATTAAAATTGACAAAGAAGTTATGGTGCCTAGT
GAAGAAGTTATTTTAAGTATGACACGTCATGGATATATTAAACGTACTTCTATTCGTAGC
TTTAATGCTAGCGGTGTTGAAGATATTGGTTTAAAAGATGGTGACAGTTTACTTAAACAT
CAAGAAGTAAATACGCAAGATACCGTACTAGTATTTACAAATAAAGGTCGTTATCTATTT
ATACCGGTTCATAAATTAGCAGATATTCGTTGGAAAGAATTGGGACAACATGTATCACAA
ATAGTTCCTATCGAAGAAGATGAAGTGGTTATTAATGTCTTTAATGAAAAGGACTTTAAT
ACAGATGCATTTTATGTTTTTGCGACTCAAAATGGCATGATTAAGAAAAGTACAGTGCCT
CTATTTAAAACAACGCGTTTTAATAAACCTTTAATTGCTACTAAAGTTAAAGAAAATGAT
GATTTGATTAGTGTTATGCGCTTTGAAAAAGATCAATTAATTACCGTCATTACTAATAAA
GGTATGTCATTAACGTATAATACAAGTGAACTATCAGATACCGGATTAAGGGCAGCTGGT
GTTAAATCAATAAATCTTAAAGCTGAAGATTTCGTTGTTATGACAGAAGGTGTTTCTGAA
AATGATACTATATTGATGGCCACACAACGCGGCTCGTTAAAACGTATTAGTTTTAAAATC
TTACAAGTTGCTAAAAGAGCACAACGTGGAATAACTTTATTAAAAGAATTAAAGAAAAAT
CCACATCGTATTGTAGCTGCACATGTAGTGACAGGTGAACATAGTCAATATACATTATAT
TCAAAATCAAATGAAGAACATGGTTTAATTAATGATATTCATAAATCTGAACAATATACA
AATGGCTCATTCATTGTAGATACAGATGATTTTGGTGAAGTAATAGACATGTATATTAGC
TAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1920
1980
2040
2100
2160
2220
2280
2340
2400
2403
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1390 [new locus tag: SACOL_RS07075 ]
- symbol: ParC
- description: DNA topoisomerase IV subunit A
- length: 800
- theoretical pI: 6.37111
- theoretical MW: 90997.4
- GRAVY: -0.382375
⊟Function[edit | edit source]
- reaction: EC 5.99.1.3? ExPASyDNA topoisomerase (ATP-hydrolyzing) ATP-dependent breakage, passage and rejoining of double-stranded DNA
- TIGRFAM: DNA metabolism DNA replication, recombination, and repair DNA topoisomerase IV, A subunit (TIGR01061; EC 5.99.1.-; HMM-score: 1054.4)DNA metabolism DNA replication, recombination, and repair DNA gyrase, A subunit (TIGR01063; EC 5.99.1.3; HMM-score: 859.8)and 2 moreDNA metabolism DNA replication, recombination, and repair DNA topoisomerase IV, A subunit (TIGR01062; EC 5.99.1.-; HMM-score: 511.5)hemerythrin-like metal-binding domain (TIGR02481; HMM-score: 13.1)
- TheSEED :
- Topoisomerase IV subunit A (EC 5.99.1.-)
DNA Metabolism DNA replication DNA topoisomerases, Type II, ATP-dependent Topoisomerase IV subunit A (EC 5.99.1.-)and 1 more - PFAM: no clan defined DNA_topoisoIV; DNA gyrase/topoisomerase IV, subunit A (PF00521; HMM-score: 532.7)and 1 moreDNA_gyraseA_C; DNA gyrase C-terminal domain, beta-propeller (PF03989; HMM-score: 88.7)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9103
- Cytoplasmic Membrane Score: 0.0021
- Cell wall & surface Score: 0.0006
- Extracellular Score: 0.087
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.003474
- TAT(Tat/SPI): 0.004737
- LIPO(Sec/SPII): 0.000528
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSEIIQDLSLEDVLGDRFGRYSKYIIQERALPDVRDGLKPVQRRILYAMYSSGNTHDKNFRKSAKTVGDVIGQYHPHGDSSVYEAMVRLSQDWKLRHVLIEMHGNNGSIDNDPPAAMRYTEAKLSLLAEELLRDINKETVSFIPNYDDTTLEPMVLPSRFPNLLVNGSTGISAGYATDIPPHNLAEVIQATLKYIDNPDITVNQLMKYIKGPDFPTGGIIQGIDGIKKAYESGKGRIIVRSKVEEETLRNGRKQLIITEIPYEVNKSSLVKRIDELRADKKVDGIVEVRDETDRTGLRIAIELKKDVNSESIKNYLYKNSDLQISYNFNMVAISDGRPKLMGIRQIIDSYLNHQIEVVANRTKFELDNAEKRMHIVEGLIKALSILDKVIELIRSSKNKRDAKENLIEVYEFTEEQAEAIVMLQLYRLTNTDIVALEGEHKELEALIKQLRHILDNHDALLNVIKEELNEIKKKFKSERLSLIEAEIEEIKIDKEVMVPSEEVILSMTRHGYIKRTSIRSFNASGVEDIGLKDGDSLLKHQEVNTQDTVLVFTNKGRYLFIPVHKLADIRWKELGQHVSQIVPIEEDEVVINVFNEKDFNTDAFYVFATQNGMIKKSTVPLFKTTRFNKPLIATKVKENDDLISVMRFEKDQLITVITNKGMSLTYNTSELSDTGLRAAGVKSINLKAEDFVVMTEGVSENDTILMATQRGSLKRISFKILQVAKRAQRGITLLKELKKNPHRIVAAHVVTGEHSQYTLYSKSNEEHGLINDIHKSEQYTNGSFIVDTDDFGEVIDMYIS
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2]
- quantitative data / protein copy number per cell: 93 [3]
- interaction partners:
SACOL2657 (arcA) arginine deiminase [4] (data from MRSA252) SACOL0557 (cysK) cysteine synthase [4] (data from MRSA252) SACOL1637 (dnaK) molecular chaperone DnaK [4] (data from MRSA252) SACOL0002 (dnaN) DNA polymerase III subunit beta [4] (data from MRSA252) SACOL0842 (eno) phosphopyruvate hydratase [4] (data from MRSA252) SACOL1016 (fabI) enoyl-ACP reductase [4] (data from MRSA252) SACOL2091 (fabZ) (3R)-hydroxymyristoyl-ACP dehydratase [4] (data from MRSA252) SACOL1329 (femC) glutamine synthetase [4] (data from MRSA252) SACOL0593 (fusA) elongation factor G [4] (data from MRSA252) SACOL1734 (gapA2) glyceraldehyde 3-phosphate dehydrogenase 2 [4] (data from MRSA252) SACOL1961 (gatA) aspartyl/glutamyl-tRNA amidotransferase subunit A [4] (data from MRSA252) SACOL1960 (gatB) aspartyl/glutamyl-tRNA amidotransferase subunit B [4] (data from MRSA252) SACOL0006 (gyrA) DNA gyrase, A subunit [4] (data from MRSA252) SACOL0005 (gyrB) DNA gyrase, B subunit [4] (data from MRSA252) SACOL1513 (hup) DNA-binding protein HU [4] (data from MRSA252) SACOL1741 (icd) isocitrate dehydrogenase [4] (data from MRSA252) SACOL0222 (ldh1) L-lactate dehydrogenase [4] (data from MRSA252) SACOL2618 (ldh2) L-lactate dehydrogenase [4] (data from MRSA252) SACOL1837 (metK) S-adenosylmethionine synthetase [4] (data from MRSA252) SACOL2092 (murAA) UDP-N-acetylglucosamine 1-carboxyvinyltransferase [4] (data from MRSA252) SACOL0746 (norR) MarR family transcriptional regulator [4] (data from MRSA252) SACOL1102 (pdhA) pyruvate dehydrogenase complex E1 component subunit alpha [4] (data from MRSA252) SACOL1105 (pdhD) dihydrolipoamide dehydrogenase [4] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [4] (data from MRSA252) SACOL0544 (prsA) ribose-phosphate pyrophosphokinase [4] (data from MRSA252) SACOL1091 (ptsH) phosphocarrier protein HPr [4] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [4] (data from MRSA252) SACOL2212 (rplQ) 50S ribosomal protein L17 [4] (data from MRSA252) SACOL1257 (rplS) 50S ribosomal protein L19 [4] (data from MRSA252) SACOL2237 (rplW) 50S ribosomal protein L23 [4] (data from MRSA252) SACOL2221 (rpmD) 50S ribosomal protein L30 [4] (data from MRSA252) SACOL0589 (rpoC) DNA-directed RNA polymerase subunit beta' [4] (data from MRSA252) SACOL1516 (rpsA) 30S ribosomal protein S1 [4] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [4] (data from MRSA252) SACOL2222 (rpsE) 30S ribosomal protein S5 [4] (data from MRSA252) SACOL0592 (rpsG) 30S ribosomal protein S7 [4] (data from MRSA252) SACOL2206 (rpsI) 30S ribosomal protein S9 [4] (data from MRSA252) SACOL0591 (rpsL) 30S ribosomal protein S12 [4] (data from MRSA252) SACOL2230 (rpsQ) 30S ribosomal protein S17 [4] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [4] (data from MRSA252) SACOL0303 5'-nucleotidase [4] (data from MRSA252) SACOL0731 LysR family transcriptional regulator [4] (data from MRSA252) SACOL0944 NADH dehydrogenase [4] (data from MRSA252) SACOL0973 fumarylacetoacetate hydrolase [4] (data from MRSA252) SACOL1759 universal stress protein [4] (data from MRSA252) SACOL2173 alkaline shock protein 23 [4] (data from MRSA252) SACOL2561 hydroxymethylglutaryl-CoA synthase [4] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: LexA (repression) regulon
LexA (TF) important in SOS response; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
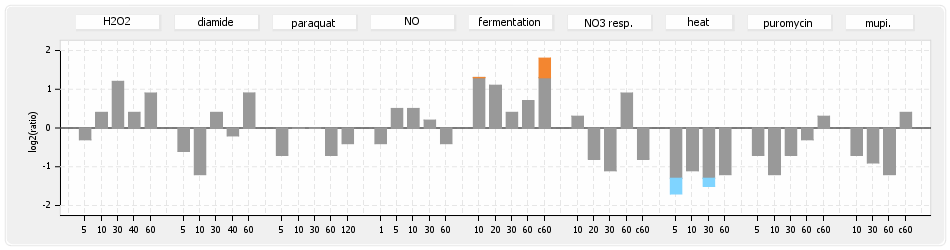
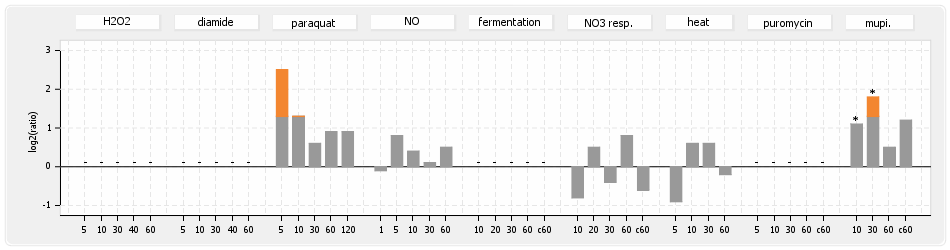
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 20.33 h [5]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 4.00 4.01 4.02 4.03 4.04 4.05 4.06 4.07 4.08 4.09 4.10 4.11 4.12 4.13 4.14 4.15 4.16 4.17 4.18 4.19 4.20 4.21 4.22 4.23 4.24 4.25 4.26 4.27 4.28 4.29 4.30 4.31 4.32 4.33 4.34 4.35 4.36 4.37 4.38 4.39 4.40 4.41 4.42 4.43 4.44 4.45 4.46 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)