Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00123
- pan locus tag?: SAUPAN000987000
- symbol: SAOUHSC_00123
- pan gene symbol?: capJ
- synonym:
- product: capsular polysaccharide biosynthesis protein Cap5J
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00123
- symbol: SAOUHSC_00123
- product: capsular polysaccharide biosynthesis protein Cap5J
- replicon: chromosome
- strand: +
- coordinates: 128497..129663
- length: 1167
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919832 NCBI
- RefSeq: YP_498723 NCBI
- BioCyc: G1I0R-114 BioCyc
- MicrobesOnline: 1288617 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141ATGAAATTTTTTGTACTTTGTGCAATTATCAGCATGAACATATTTATAGTAATCTCTACA
TTTACTAAAGAAGTATTAGGGTTCCCTATAGAGCCGGTGTATTACTCAACCATGGTTGGT
ATAGCATTAATTACTACGGTGTTTGCTATTTATAAGATAATTGTCACCCAAGAAATTCCG
CGAGGGTTAATATTATTAATTGCTATATGTTTGCTTTATCTAGCTTTTTATTATTTTTCA
CCAGATAAGGAAGAGAAACTAGCTAAAAATAATATTCTATTCTTTTTAACATGGGCAGTT
CCAGCGGCAATTAGTGGTATTTATATTAAATATATAAACAAGGCTACGGTAGAAAGATTT
TTTAAATTAGTATTTTTCATATTTTCTGTTTCATTTATTTTTGTAATTTTAATACCAAAA
CTTACAGGTGAGATACCTAGCTATATCAATTTTGGACTTATGAACTATCAAAACGCTTCG
TACCTTTCAGCATTTACTGCCGGATTAGGCATTTATTTCATTATGAAAGGTTCAGTTAAA
CATAAGTGGATATATGTTCTATTTACAATAATTGATATCCCTATTGTGTTTATACCAGGA
GGGCGTGGAGGTGCTATTTTATTAATTCTTTACGGCTTATTTGCATTTATACTTATTACG
TTTAAAAGAGGAATACCTATCGCAGTAAAAAGCATTATGTATATTTTTGCATTAAGCATA
TCTAGTGTATTGATTTACTTTCTTTTTACAAAAGGTTCGAATACTAGAACATTTTCATAT
CTACAAGGTGGAACACTTAATTTAGAAGGTACTTCTGGAAGAGGACCGATTTATGAAAAA
GGTATTTACTTTATTCAACAAAGTTCGTTATTAGGCTATGGGCCATTTAACTATTATAAA
CTAATCGGAAATATACCACATAACATCATTATTGAGTTGATTCTATCATTTGGCTTATTA
GGGTTTTTTATCATAATGATTTGCATTTTGCTACTAGTTTATAAAATGATTAGGAACTAT
GATCCAAACACTATAGATTTACTCGTTATGTTTATAGCAATCTATCCAATCACATTATTA
ATGTTTAGTTCAAATTATTTAGTTGTAAGTGAATTTTGGTTTGTGTTGTTCTATTTTATT
ACAAAAGGACGGCGTCATCATGGCTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1167
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00123
- symbol: SAOUHSC_00123
- description: capsular polysaccharide biosynthesis protein Cap5J
- length: 388
- theoretical pI: 9.76373
- theoretical MW: 44259
- GRAVY: 0.930928
⊟Function[edit | edit source]
- TIGRFAM: Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides oligosaccharide repeat unit polymerase (TIGR04370; HMM-score: 17.9)
- TheSEED :
- Capsular polysaccharide synthesis enzyme Cap5J
- PFAM: O-anti_assembly (CL0499) Wzy_C; O-Antigen ligase (PF04932; HMM-score: 32.7)and 1 moreno clan defined PrgI; PrgI family protein (PF12666; HMM-score: 12.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 11
- DeepLocPro: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 0.9956
- Cell wall & surface Score: 0
- Extracellular Score: 0.0044
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0
- Signal peptide possibility: 0
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.249095
- TAT(Tat/SPI): 0.000307
- LIPO(Sec/SPII): 0.050797
- predicted transmembrane helices (TMHMM): 11
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKFFVLCAIISMNIFIVISTFTKEVLGFPIEPVYYSTMVGIALITTVFAIYKIIVTQEIPRGLILLIAICLLYLAFYYFSPDKEEKLAKNNILFFLTWAVPAAISGIYIKYINKATVERFFKLVFFIFSVSFIFVILIPKLTGEIPSYINFGLMNYQNASYLSAFTAGLGIYFIMKGSVKHKWIYVLFTIIDIPIVFIPGGRGGAILLILYGLFAFILITFKRGIPIAVKSIMYIFALSISSVLIYFLFTKGSNTRTFSYLQGGTLNLEGTSGRGPIYEKGIYFIQQSSLLGYGPFNYYKLIGNIPHNIIIELILSFGLLGFFIIMICILLLVYKMIRNYDPNTIDLLVMFIAIYPITLLMFSSNYLVVSEFWFVLFYFITKGRRHHG
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [1] : S37 > SAOUHSC_00114 > SAOUHSC_00115 > SAOUHSC_00116 > SAOUHSC_00117 > SAOUHSC_00118 > SAOUHSC_00119 > SAOUHSC_00120 > SAOUHSC_00121 > SAOUHSC_00122 > SAOUHSC_00123 > SAOUHSC_00124 > SAOUHSC_00125 > SAOUHSC_00126 > SAOUHSC_00127predicted SigB promoter [1] : SAOUHSC_00114 > SAOUHSC_00115 > SAOUHSC_00116 > SAOUHSC_00117 > SAOUHSC_00118 > SAOUHSC_00119 > SAOUHSC_00120 > SAOUHSC_00121 > SAOUHSC_00122 > SAOUHSC_00123 > SAOUHSC_00124 > SAOUHSC_00125 > SAOUHSC_00126 > SAOUHSC_00127 > SAOUHSC_00128 > S38 > SAOUHSC_00129
⊟Regulation[edit | edit source]
- regulators: CodY* (repression) regulon, SigB* (activation) regulon
CodY* (TF) important in Amino acid metabolism; RegPrecise transcription unit transferred from N315 data RegPrecise SigB* (sigma factor) controlling a large regulon involved in stress/starvation response and adaptation; [4] [1] other strains
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
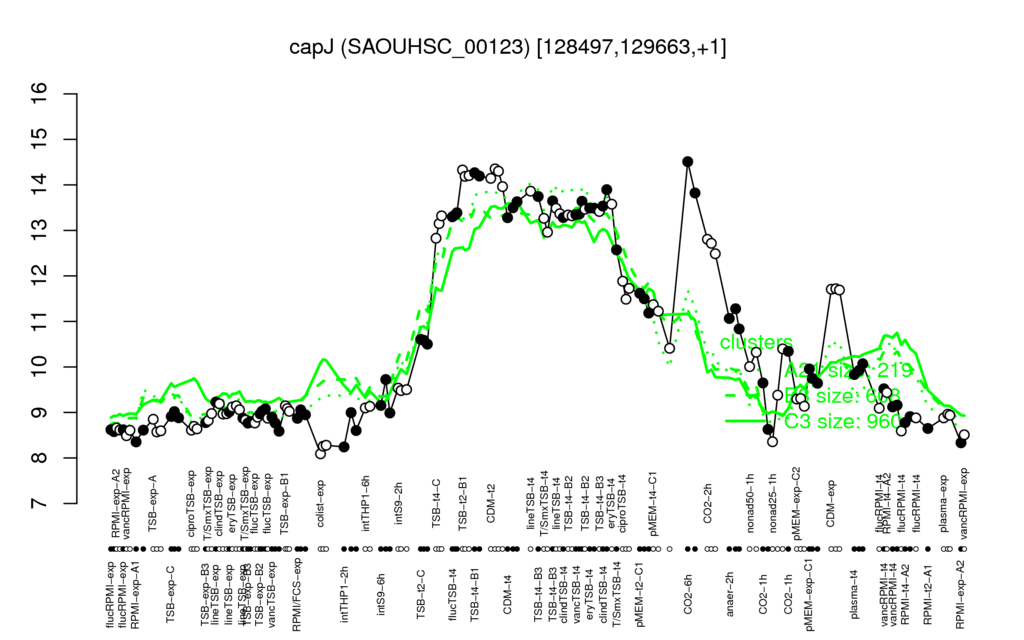 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ 1.0 1.1 1.2 1.3 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e) - ↑ Daniela Keinhörster, Shilpa Elizabeth George, Christopher Weidenmaier, Christiane Wolz
Function and regulation of Staphylococcus aureus wall teichoic acids and capsular polysaccharides.
Int J Med Microbiol: 2019, 309(6);151333
[PubMed:31362856] [WorldCat.org] [DOI] (I p) - ↑ S Sau, J Sun, C Y Lee
Molecular characterization and transcriptional analysis of type 8 capsule genes in Staphylococcus aureus.
J Bacteriol: 1997, 179(5);1614-21
[PubMed:9045821] [WorldCat.org] [DOI] (P p) - ↑ Markus Bischoff, Paul Dunman, Jan Kormanec, Daphne Macapagal, Ellen Murphy, William Mounts, Brigitte Berger-Bächi, Steven Projan
Microarray-based analysis of the Staphylococcus aureus sigmaB regulon.
J Bacteriol: 2004, 186(13);4085-99
[PubMed:15205410] [WorldCat.org] [DOI] (P p)
