Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00847
- pan locus tag?: SAUPAN002894000
- symbol: SAOUHSC_00847
- pan gene symbol?: sufC
- synonym:
- product: ABC transporter ATP-binding protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
⊟Accession numbers[edit | edit source]
- Gene ID: 3918959 NCBI
- RefSeq: YP_499401 NCBI
- BioCyc: G1I0R-791 BioCyc
- MicrobesOnline: 1289312 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721ATGGCATCAACATTAGAAATCAAAGACCTACATGTGTCTATTGAGGATAAAGAAATCTTA
AAAGGTGTTAACTTGACAATTAACACTGATGAAATACATGCGATTATGGGACCAAACGGG
ACAGGTAAATCAACTTTATCATCTGCAATTATGGGACACCCAAGCTATGAAGTAACTAAA
GGAGAAGTACTTTTAGACGGTGTAAATATTTTAGAATTAGAAGTTGATGAAAGAGCAAAA
GCAGGATTATTCTTGGCAATGCAATATCCATCAGAAATTACAGGTGTTACAAATGCTGAT
TTCATGCGTTCAGCAATCAATGCGAAACGTGAAGAAGGACAAGAAATCAACTTAATGCAA
TTTATTAAGAAATTAGATAAAAACATGGATTTTCTAGACATAGATAAAGACATGGCACAA
CGTTATTTAAATGAAGGTTTCTCAGGTGGAGAGAAGAAACGTAACGAAATCTTACAATTA
ATGATGTTAGAACCTAAGTTTGCAATCTTAGATGAAATCGATTCAGGGTTAGACATCGAT
GCATTAAAAGTTGTATCTAAAGGTATTAACCAAATGCGTGGGGAAAACTTTGGTGCATTA
ATGATTACACACTATCAACGATTATTAAATTACATTACTCCTGATAAAGTACATGTAATG
TATGCTGGTAAAGTCGTTAAATCTGGTGGTCCAGAATTAGCAAAACGTCTTGAAGAAGAA
GGATATGAATGGGTTAAAGAAGAGTTCGGTTCAGCTGAATAA60
120
180
240
300
360
420
480
540
600
660
720
762
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00847
- symbol: SAOUHSC_00847
- description: ABC transporter ATP-binding protein
- length: 253
- theoretical pI: 4.61058
- theoretical MW: 28275.2
- GRAVY: -0.303162
⊟Function[edit | edit source]
- TIGRFAM: Biosynthesis of cofactors, prosthetic groups, and carriers Other FeS assembly ATPase SufC (TIGR01978; HMM-score: 369.2)and 73 moreCell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides LPS export ABC transporter ATP-binding protein (TIGR04406; HMM-score: 104)Transport and binding proteins Other LPS export ABC transporter ATP-binding protein (TIGR04406; HMM-score: 104)Transport and binding proteins Amino acids, peptides and amines urea ABC transporter, ATP-binding protein UrtE (TIGR03410; HMM-score: 83.7)Transport and binding proteins Amino acids, peptides and amines urea ABC transporter, ATP-binding protein UrtD (TIGR03411; HMM-score: 81.4)Transport and binding proteins Unknown substrate energy-coupling factor transporter ATPase (TIGR04520; EC 3.6.3.-; HMM-score: 81.3)Transport and binding proteins Other daunorubicin resistance ABC transporter, ATP-binding protein (TIGR01188; HMM-score: 74.1)Transport and binding proteins Unknown substrate energy-coupling factor transporter ATPase (TIGR04521; EC 3.6.3.-; HMM-score: 73.9)Transport and binding proteins Cations and iron carrying compounds nickel import ATP-binding protein NikE (TIGR02769; EC 3.6.3.24; HMM-score: 72.9)Transport and binding proteins Anions phosphate ABC transporter, ATP-binding protein (TIGR00972; EC 3.6.3.27; HMM-score: 70.9)ABC transporter, permease/ATP-binding protein (TIGR02204; HMM-score: 69.1)Transport and binding proteins Carbohydrates, organic alcohols, and acids ABC transporter, ATP-binding subunit, PQQ-dependent alcohol dehydrogenase system (TIGR03864; HMM-score: 69)Cellular processes Pathogenesis type I secretion system ATPase (TIGR03375; HMM-score: 67.8)Protein fate Protein and peptide secretion and trafficking type I secretion system ATPase (TIGR03375; HMM-score: 67.8)proposed F420-0 ABC transporter, ATP-binding protein (TIGR03873; HMM-score: 66.5)Protein fate Protein and peptide secretion and trafficking type I secretion system ATPase (TIGR01842; HMM-score: 65.6)Transport and binding proteins Amino acids, peptides and amines putative 2-aminoethylphosphonate ABC transporter, ATP-binding protein (TIGR03265; HMM-score: 65.3)thiol reductant ABC exporter, CydD subunit (TIGR02857; HMM-score: 65)Transport and binding proteins Cations and iron carrying compounds nickel import ATP-binding protein NikD (TIGR02770; HMM-score: 63.3)Transport and binding proteins Cations and iron carrying compounds cobalt ABC transporter, ATP-binding protein (TIGR01166; HMM-score: 61.8)Transport and binding proteins Anions sulfate ABC transporter, ATP-binding protein (TIGR00968; EC 3.6.3.25; HMM-score: 59.9)Energy metabolism Methanogenesis methyl coenzyme M reductase system, component A2 (TIGR03269; HMM-score: 59)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides lipid A export permease/ATP-binding protein MsbA (TIGR02203; HMM-score: 58.9)Transport and binding proteins Other lipid A export permease/ATP-binding protein MsbA (TIGR02203; HMM-score: 58.9)Transport and binding proteins Other antigen peptide transporter 2 (TIGR00958; HMM-score: 57.4)Transport and binding proteins Other pigment precursor permease (TIGR00955; HMM-score: 56.5)Cellular processes Biosynthesis of natural products NHLM bacteriocin system ABC transporter, peptidase/ATP-binding protein (TIGR03796; HMM-score: 56.2)Transport and binding proteins Amino acids, peptides and amines NHLM bacteriocin system ABC transporter, peptidase/ATP-binding protein (TIGR03796; HMM-score: 56.2)Cellular processes Biosynthesis of natural products NHLM bacteriocin system ABC transporter, ATP-binding protein (TIGR03797; HMM-score: 56.2)Transport and binding proteins Amino acids, peptides and amines NHLM bacteriocin system ABC transporter, ATP-binding protein (TIGR03797; HMM-score: 56.2)2-aminoethylphosphonate ABC transport system, ATP-binding component PhnT (TIGR03258; HMM-score: 55.6)Cellular processes Toxin production and resistance putative bacteriocin export ABC transporter, lactococcin 972 group (TIGR03608; HMM-score: 55.4)Transport and binding proteins Unknown substrate putative bacteriocin export ABC transporter, lactococcin 972 group (TIGR03608; HMM-score: 55.4)thiol reductant ABC exporter, CydC subunit (TIGR02868; HMM-score: 52.8)Protein fate Protein and peptide secretion and trafficking ABC-type bacteriocin transporter (TIGR01193; HMM-score: 51.2)Protein fate Protein modification and repair ABC-type bacteriocin transporter (TIGR01193; HMM-score: 51.2)Transport and binding proteins Other ABC-type bacteriocin transporter (TIGR01193; HMM-score: 51.2)lantibiotic protection ABC transporter, ATP-binding subunit (TIGR03740; HMM-score: 50.9)Transport and binding proteins Anions phosphonate ABC transporter, ATP-binding protein (TIGR02315; EC 3.6.3.28; HMM-score: 50.3)Protein fate Protein and peptide secretion and trafficking lipoprotein releasing system, ATP-binding protein (TIGR02211; EC 3.6.3.-; HMM-score: 49.9)ATP-binding cassette protein, ChvD family (TIGR03719; HMM-score: 49.2)Protein fate Protein and peptide secretion and trafficking type I secretion system ATPase (TIGR01846; HMM-score: 49)ABC exporter ATP-binding subunit, DevA family (TIGR02982; HMM-score: 48.7)ectoine/hydroxyectoine ABC transporter, ATP-binding protein EhuA (TIGR03005; HMM-score: 47.9)Transport and binding proteins Carbohydrates, organic alcohols, and acids D-xylose ABC transporter, ATP-binding protein (TIGR02633; EC 3.6.3.17; HMM-score: 47.4)Protein fate Protein and peptide secretion and trafficking heme ABC exporter, ATP-binding protein CcmA (TIGR01189; EC 3.6.3.41; HMM-score: 47.3)Transport and binding proteins Other heme ABC exporter, ATP-binding protein CcmA (TIGR01189; EC 3.6.3.41; HMM-score: 47.3)Transport and binding proteins Amino acids, peptides and amines glycine betaine/L-proline transport ATP binding subunit (TIGR01186; HMM-score: 45.3)Transport and binding proteins Other thiamine ABC transporter, ATP-binding protein (TIGR01277; EC 3.6.3.-; HMM-score: 43.3)Cellular processes Other nodulation ABC transporter NodI (TIGR01288; HMM-score: 42.3)Transport and binding proteins Other nodulation ABC transporter NodI (TIGR01288; HMM-score: 42.3)Transport and binding proteins Carbohydrates, organic alcohols, and acids glucan exporter ATP-binding protein (TIGR01192; EC 3.6.3.42; HMM-score: 41.7)D-methionine ABC transporter, ATP-binding protein (TIGR02314; EC 3.6.3.-; HMM-score: 41.1)Cellular processes Cell division cell division ATP-binding protein FtsE (TIGR02673; HMM-score: 41)Transport and binding proteins Anions nitrate ABC transporter, ATP-binding proteins C and D (TIGR01184; HMM-score: 40.2)Transport and binding proteins Other nitrate ABC transporter, ATP-binding proteins C and D (TIGR01184; HMM-score: 40.2)Transport and binding proteins Other pleiotropic drug resistance family protein (TIGR00956; HMM-score: 39.9)Transport and binding proteins Unknown substrate anchored repeat-type ABC transporter, ATP-binding subunit (TIGR03771; HMM-score: 39.5)phosphonate C-P lyase system protein PhnL (TIGR02324; HMM-score: 39.2)Transport and binding proteins Amino acids, peptides and amines polyamine ABC transporter, ATP-binding protein (TIGR01187; EC 3.6.3.31; HMM-score: 38)gliding motility-associated ABC transporter ATP-binding subunit GldA (TIGR03522; HMM-score: 37.5)Transport and binding proteins Anions molybdate ABC transporter, ATP-binding protein (TIGR02142; EC 3.6.3.29; HMM-score: 36.3)Transport and binding proteins Carbohydrates, organic alcohols, and acids peroxysomal long chain fatty acyl transporter (TIGR00954; HMM-score: 29.4)Central intermediary metabolism Phosphorus compounds phosphonate C-P lyase system protein PhnK (TIGR02323; HMM-score: 29.4)Transport and binding proteins Amino acids, peptides and amines cyclic peptide transporter (TIGR01194; HMM-score: 26)Transport and binding proteins Other cyclic peptide transporter (TIGR01194; HMM-score: 26)Transport and binding proteins Other multi drug resistance-associated protein (MRP) (TIGR00957; HMM-score: 24.3)Transport and binding proteins Other rim ABC transporter (TIGR01257; HMM-score: 19.4)Transport and binding proteins Amino acids, peptides and amines choline ABC transporter, ATP-binding protein (TIGR03415; HMM-score: 19.2)Transport and binding proteins Anions cystic fibrosis transmembrane conductor regulator (CFTR) (TIGR01271; EC 3.6.3.49; HMM-score: 16.7)Cellular processes Cell division chromosome segregation protein SMC (TIGR02168; HMM-score: 15.3)DNA metabolism Chromosome-associated proteins chromosome segregation protein SMC (TIGR02168; HMM-score: 15.3)carbohydrate kinase, thermoresistant glucokinase family (TIGR01313; EC 2.7.1.-; HMM-score: 12.7)DNA metabolism DNA replication, recombination, and repair excinuclease ABC subunit A (TIGR00630; EC 3.1.25.-; HMM-score: 12.2)
- TheSEED :
- Iron-sulfur cluster assembly ATPase protein SufC
- PFAM: P-loop_NTPase (CL0023) ABC_tran; ABC transporter (PF00005; HMM-score: 81.8)and 21 moreSMC_N; RecF/RecN/SMC N terminal domain (PF02463; HMM-score: 35.9)Aha1_BPI (CL0648) DUF1439; Protein of unknown function (DUF1439) (PF07273; HMM-score: 20.3)P-loop_NTPase (CL0023) AAA_29; P-loop containing region of AAA domain (PF13555; HMM-score: 20)AAA_28; AAA domain (PF13521; HMM-score: 19.8)RsgA_GTPase; RsgA GTPase (PF03193; HMM-score: 17.3)AAA_27; AAA domain (PF13514; HMM-score: 16.7)AAA_21; AAA domain, putative AbiEii toxin, Type IV TA system (PF13304; HMM-score: 16.4)AAA_15; AAA ATPase domain (PF13175; HMM-score: 16.2)AAA_18; AAA domain (PF13238; HMM-score: 16.2)AAA_14; AAA domain (PF13173; HMM-score: 16.1)AAA; ATPase family associated with various cellular activities (AAA) (PF00004; HMM-score: 15.2)AAA_23; AAA domain (PF13476; HMM-score: 15)AAA_13; AAA domain (PF13166; HMM-score: 14.3)AAA_33; AAA domain (PF13671; HMM-score: 14)no clan defined PFam54_60; Borrelia Bbcrasp-1 domain containing protein (PF05714; HMM-score: 13.6)P-loop_NTPase (CL0023) AAA_25; AAA domain (PF13481; HMM-score: 13.5)GT-B (CL0113) Glyco_trans_4_2; Glycosyl transferase 4-like (PF13477; HMM-score: 13.4)P-loop_NTPase (CL0023) AAA_22; AAA domain (PF13401; HMM-score: 13.3)AAA_5; AAA domain (dynein-related subfamily) (PF07728; HMM-score: 13.2)Adhesin (CL0204) Sgo0707_N2; Sgo0707 N2 domain (PF20623; HMM-score: 12)no clan defined DUF1420; Protein of unknown function (DUF1420) (PF07220; HMM-score: 10.1)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.73
- Cytoplasmic Membrane Score: 0.2395
- Cell wall & surface Score: 0
- Extracellular Score: 0.0304
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.007752
- TAT(Tat/SPI): 0.000871
- LIPO(Sec/SPII): 0.000474
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MASTLEIKDLHVSIEDKEILKGVNLTINTDEIHAIMGPNGTGKSTLSSAIMGHPSYEVTKGEVLLDGVNILELEVDERAKAGLFLAMQYPSEITGVTNADFMRSAINAKREEGQEINLMQFIKKLDKNMDFLDIDKDMAQRYLNEGFSGGEKKRNEILQLMMLEPKFAILDEIDSGLDIDALKVVSKGINQMRGENFGALMITHYQRLLNYITPDKVHVMYAGKVVKSGGPELAKRLEEEGYEWVKEEFGSAE
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [2] [3]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_00847 > SAOUHSC_00848 > SAOUHSC_00849 > SAOUHSC_00850 > SAOUHSC_00851predicted SigB promoter [4] : SAOUHSC_00845 > S351 > S353 > SAOUHSC_00847 > S354 > SAOUHSC_00848 > S355 > SAOUHSC_00849 > SAOUHSC_00850 > S356 > SAOUHSC_00851 > S357predicted SigB promoter [4] : S353 > SAOUHSC_00847 > S354 > SAOUHSC_00848 > S355 > SAOUHSC_00849 > SAOUHSC_00850 > S356 > SAOUHSC_00851 > S357
⊟Regulation[edit | edit source]
- regulators: PerR* (repression) regulon, SigB* (activation) regulon
PerR* (TF) important in Oxidative stress response; RegPrecise SigB* (sigma factor) controlling a large regulon involved in stress/starvation response and adaptation; [4] other strains
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
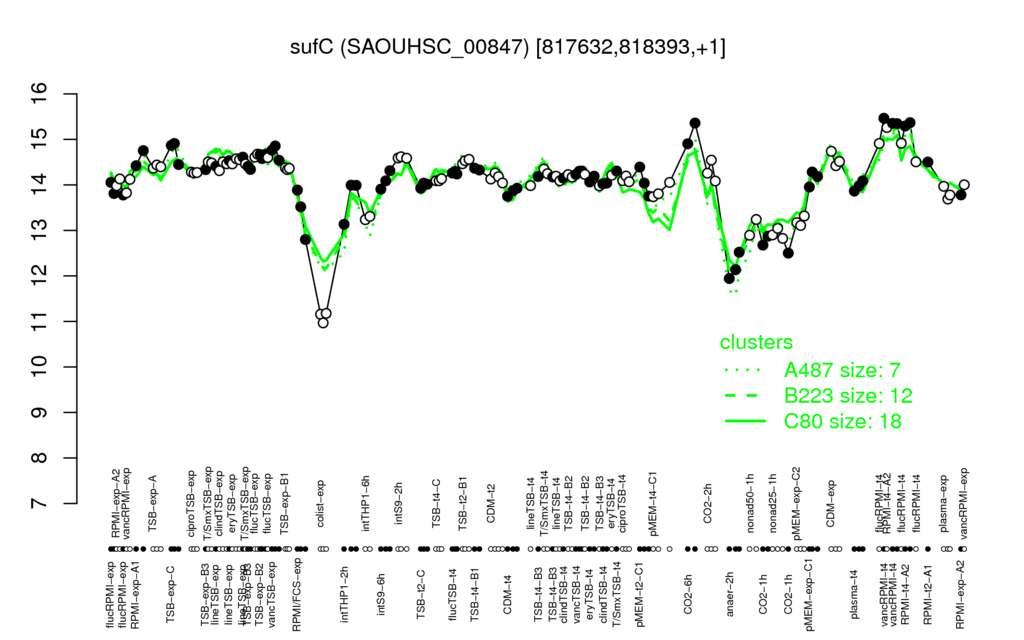 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Roy R Chaudhuri, Andrew G Allen, Paul J Owen, Gil Shalom, Karl Stone, Marcus Harrison, Timothy A Burgis, Michael Lockyer, Jorge Garcia-Lara, Simon J Foster, Stephen J Pleasance, Sarah E Peters, Duncan J Maskell, Ian G Charles
Comprehensive identification of essential Staphylococcus aureus genes using Transposon-Mediated Differential Hybridisation (TMDH).
BMC Genomics: 2009, 10;291
[PubMed:19570206] [WorldCat.org] [DOI] (I e) - ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 4.0 4.1 4.2 4.3 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
