Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL2488 [new locus tag: SACOL_RS13040 ]
- pan locus tag?: SAUPAN006056000
- symbol: SACOL2488
- pan gene symbol?: —
- synonym:
- product: short chain dehydrogenase/reductase oxidoreductase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL2488 [new locus tag: SACOL_RS13040 ]
- symbol: SACOL2488
- product: short chain dehydrogenase/reductase oxidoreductase
- replicon: chromosome
- strand: -
- coordinates: 2543619..2544314
- length: 696
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238118 NCBI
- RefSeq: YP_187284 NCBI
- BioCyc: see SACOL_RS13040
- MicrobesOnline: 913962 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661ATGACAGTATTAACAGATAAAGTAGCAGTAGTTACAGGTGCAGGTAGTGGTATTGGAGAA
GCAATTGCAACATTACTACATGAAGAAGGGGCAAAAGTTGTCTTAGCAGGTAGAAATAAA
GAAAAATTACAAAACGTAGCGAATCAATTGTCACAAGATAGTGTGAAGGTAGTGCCAACA
GATGTAACGAATAAAGAAGAAGTCGATGAATTGATAAAAATTGCACAACAAACATTCGGT
GGTTTGGATATTGTTATCAATAGTGCGGGGCAAATGTTGTCGTCTAAGATTACTGATTAT
CAAGTAGATGAGTGGGATAGTATGATTGATGTGAATATCAAAGGCACTTTATATACGGCA
CAGGCTGCATTACCAACTATGTTAGAACAATCAAGTGGCCATCTTATTAACATTGCATCT
ATTTCTGGCTTTGAAGTAACGAAAAGTAGTACGATTTATAGTGCGACGAAAGCAGCAGTT
CACACTATTACTCAAGGATTAGAAAAAGAGTTGGCAAAGACAGGCGTTAAAGTAACAAGC
ATTTCTCCAGGAATGGTAGATACAGCCATAACTGCCGCATACAATCCAACAGATCGTAAA
AAACTTGAACCACAAGATATTGCAGAAGCAGTATTATATGCATTAACACAACCAAAGCAC
GTTAACGTGAATGAAATTACAGTGCGTCCTGTATAA60
120
180
240
300
360
420
480
540
600
660
696
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL2488 [new locus tag: SACOL_RS13040 ]
- symbol: SACOL2488
- description: short chain dehydrogenase/reductase oxidoreductase
- length: 231
- theoretical pI: 4.91397
- theoretical MW: 24603.9
- GRAVY: -0.0290043
⊟Function[edit | edit source]
- reaction: EC 1.-.-.-? ExPASy
- TIGRFAM: 3-hydroxybutyrate dehydrogenase (TIGR01963; HMM-score: 155.3)Energy metabolism Fermentation acetoin reductases (TIGR02415; EC 1.1.1.-; HMM-score: 149.3)Fatty acid and phospholipid metabolism Biosynthesis 3-oxoacyl-[acyl-carrier-protein] reductase (TIGR01830; EC 1.1.1.100; HMM-score: 147.8)acetoacetyl-CoA reductase (TIGR01829; EC 1.1.1.36; HMM-score: 126.4)Unknown function Enzymes of unknown specificity SDR family mycofactocin-dependent oxidoreductase (TIGR03971; EC 1.1.99.-; HMM-score: 125.5)rhamnulose-1-phosphate aldolase/alcohol dehydrogenase (TIGR02632; EC 1.1.1.1,4.1.2.19; HMM-score: 124.7)and 18 more2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase (TIGR04316; EC 1.3.1.28; HMM-score: 123.5)2-hydroxycyclohexanecarboxyl-CoA dehydrogenase (TIGR03206; EC 1.1.1.-; HMM-score: 119.2)Energy metabolism Biosynthesis and degradation of polysaccharides 2-deoxy-D-gluconate 3-dehydrogenase (TIGR01832; EC 1.1.1.125; HMM-score: 117)Unknown function Enzymes of unknown specificity SDR family mycofactocin-dependent oxidoreductase (TIGR04504; EC 1.1.99.-; HMM-score: 114.8)cis-2,3-dihydrobiphenyl-2,3-diol dehydrogenase (TIGR03325; EC 1.3.1.56; HMM-score: 83.7)Fatty acid and phospholipid metabolism Biosynthesis putative 3-oxoacyl-(acyl-carrier-protein) reductase (TIGR01831; HMM-score: 80)sepiapterin reductase (TIGR01500; EC 1.1.1.153; HMM-score: 58.3)pteridine reductase (TIGR02685; EC 1.5.1.33; HMM-score: 46.3)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll light-dependent protochlorophyllide reductase (TIGR01289; EC 1.3.1.33; HMM-score: 34.9)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin glutamyl-tRNA reductase (TIGR01035; EC 1.2.1.70; HMM-score: 21)Hypothetical proteins Conserved TIGR01777 family protein (TIGR01777; HMM-score: 20)Amino acid biosynthesis Aromatic amino acid family shikimate dehydrogenase (TIGR00507; EC 1.1.1.25; HMM-score: 19.9)hopanoid-associated sugar epimerase (TIGR03466; HMM-score: 17.1)Mobile and extrachromosomal element functions Plasmid functions integrating conjugative element protein PilL, PFGI-1 class (TIGR03748; HMM-score: 16.4)Unknown function Enzymes of unknown specificity putative NAD(P)H quinone oxidoreductase, PIG3 family (TIGR02824; HMM-score: 15.8)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides dTDP-glucose 4,6-dehydratase (TIGR01181; EC 4.2.1.46; HMM-score: 11.3)Protein synthesis Ribosomal proteins: synthesis and modification ribosomal protein uS11 (TIGR03632; HMM-score: 11.1)Fatty acid and phospholipid metabolism Biosynthesis hopanoid C-3 methylase HpnR (TIGR04367; HMM-score: 10.8)
- TheSEED :
- Oxidoreductase, short-chain dehydrogenase/reductase family
- PFAM: NADP_Rossmann (CL0063) adh_short; short chain dehydrogenase (PF00106; HMM-score: 196.2)adh_short_C2; Enoyl-(Acyl carrier protein) reductase (PF13561; HMM-score: 159.8)and 25 moreKR; KR domain (PF08659; HMM-score: 62.7)NAD_binding_10; NAD(P)H-binding (PF13460; HMM-score: 41.6)Sacchrp_dh_NADP; Saccharopine dehydrogenase NADP binding domain (PF03435; HMM-score: 38.1)Epimerase; NAD dependent epimerase/dehydratase family (PF01370; HMM-score: 35.1)Shikimate_DH; Shikimate / quinate 5-dehydrogenase (PF01488; HMM-score: 26.4)3HCDH_N; 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain (PF02737; HMM-score: 24.6)Polysacc_synt_2; Polysaccharide biosynthesis protein (PF02719; HMM-score: 24)SDR; SDR-like rossmann domain (PF23441; HMM-score: 23.4)F420_oxidored; NADP oxidoreductase coenzyme F420-dependent (PF03807; HMM-score: 21.4)GDP_Man_Dehyd; GDP-mannose 4,6 dehydratase (PF16363; HMM-score: 19.7)ApbA; Ketopantoate reductase PanE/ApbA (PF02558; HMM-score: 19.5)NAD_Gly3P_dh_N; NAD-dependent glycerol-3-phosphate dehydrogenase N-terminus (PF01210; HMM-score: 18.8)NmrA; NmrA-like family (PF05368; HMM-score: 17.8)S11_L18p (CL0267) Ribosomal_S11; Ribosomal protein S11 (PF00411; HMM-score: 17.4)no clan defined Arg_tRNA_synt_N; Arginyl tRNA synthetase N terminal domain (PF03485; HMM-score: 16.5)NADP_Rossmann (CL0063) ADH_zinc_N; Zinc-binding dehydrogenase (PF00107; HMM-score: 15.8)no clan defined Asp23; Asp23 family, cell envelope-related function (PF03780; HMM-score: 15)NADP_Rossmann (CL0063) THF_DHG_CYH_C; Tetrahydrofolate dehydrogenase/cyclohydrolase, NAD(P)-binding domain (PF02882; HMM-score: 14.8)TrkA_N; TrkA-N domain (PF02254; HMM-score: 14.1)RNase_H (CL0219) DDE_Tnp_1_3; Transposase DDE domain (PF13612; HMM-score: 13.9)NADP_Rossmann (CL0063) KARI_N; Acetohydroxy acid isomeroreductase, NADPH-binding domain (PF07991; HMM-score: 12.9)DUF1776; Fungal family of unknown function (DUF1776) (PF08643; HMM-score: 12.9)P-loop_NTPase (CL0023) Helicase_RecD; Helicase (PF05127; HMM-score: 12.7)NADP_Rossmann (CL0063) YjeF_N; YjeF-related protein N-terminus (PF03853; HMM-score: 11.7)no clan defined Nitro_FeMo-Co; Dinitrogenase iron-molybdenum cofactor (PF02579; HMM-score: 11.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.67
- Cytoplasmic Membrane Score: 0.01
- Cellwall Score: 0.15
- Extracellular Score: 0.17
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9289
- Cytoplasmic Membrane Score: 0.0423
- Cell wall & surface Score: 0.0053
- Extracellular Score: 0.0235
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.009386
- TAT(Tat/SPI): 0.001928
- LIPO(Sec/SPII): 0.00181
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTVLTDKVAVVTGAGSGIGEAIATLLHEEGAKVVLAGRNKEKLQNVANQLSQDSVKVVPTDVTNKEEVDELIKIAQQTFGGLDIVINSAGQMLSSKITDYQVDEWDSMIDVNIKGTLYTAQAALPTMLEQSSGHLINIASISGFEVTKSSTIYSATKAAVHTITQGLEKELAKTGVKVTSISPGMVDTAITAAYNPTDRKKLEPQDIAEAVLYALTQPKHVNVNEITVRPV
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3]
- quantitative data / protein copy number per cell: 932 [4]
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
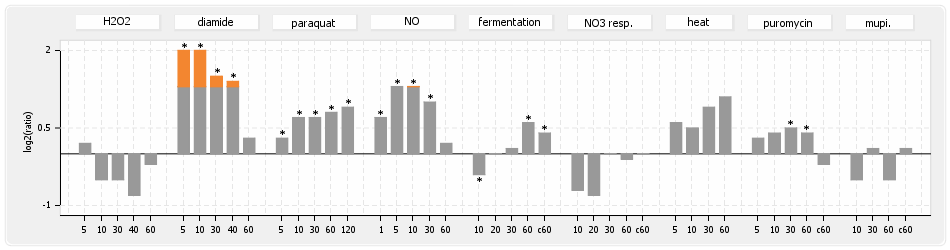
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 32.05 h [5]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)