Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00973
- pan locus tag?: SAUPAN003234000
- symbol: SAOUHSC_00973
- pan gene symbol?: tarM
- synonym:
- product:
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
⊟Accession numbers[edit | edit source]
- Gene ID: 3920118 NCBI
- RefSeq:
- BioCyc: G1I0R-917 BioCyc
- MicrobesOnline: 1289439 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441ATGAAAAAAATATTTATGATGGTACATGAGTTAGATGTGAATAAAGGTGGTATGACCTCT
TCGATGTTCAATAGAAGTAAAGAGTTTTATGATGCGGACATACCTGCTGATATTGTTACT
TTCGATTACAAAGGAAACTATGATGAAATTATTAAAGCTTTGAAAAAACAAGGTAAAATG
GATCGAAGAACGAAAATGTATAATGTATTTGAGTATTTTAAACAAATTTCAAATAATAAA
CATTTTAAGTCTAATAAATTGTTATATAAACATATTTCAGAAAGACTAAAAAATACGATT
GAAATTGAAGAGAGTAAAGGTATTTCAAGATATTTTGATATAACGACTGGTACATATATT
GCCTACATTAGAAAAAGTAAATCTGAAAAAGTGATTGATTTCTTTAAAGATAATAAACGA
ATTGAACGGTTTAGTTTTATAGATAATAAAGTGCATATGAAGGAAACATTTAATGTAGAT
AATAAAGTTTGTTATCAAGTATTTTATGATGAAAAGGGATACCCATATATTTCAAGGAAT
ATTAATGCTAATAATGGTGCTGTAGGTAAAACTTATGTGTTAGTTAATAAAAAAGAATTT
AAAAACAATTTAGCACTGTGTGTTTACTATTTAGAAAAACTAATAAAAGATTCTAAAGAT
AGTATTATGATTTGTGATGGACCAGGGAGTTTTCCAAAAATGTTTAATACAAATCATAAA
AATGCTCAGAAATATGGCGTTATTCATGTTAATCATCATGAAAATTTCGATGATACGGGT
GCATTTAAAAAAAGTGAGAAATATATTATTGAGAATGCGAATAAAATTAACGGTGTAATT
GTATTAACAGAGGCACAAAGATTAGATATTCTTAATCAATTTGATGTAGAAAATATTTTC
ACTATTAGCAATTTTGTTAAGATACATAATGCTCCAAAACATTTTCAAACTGAAAAAATC
GTAGGTCATATTTCTAGAATGGTACCAACGAAGCGAATTGATTTGCTTATTGAAGTGGCT
GAGTTAGTCGTAAAAAAAGATAATGCTGTTAAATTTCATATATATGGAGAAGGATCTGTC
AAAGATAAAATAGCTAAAATGATTGAAGATAAAAATTTAGAAAGAAATGTTTTTCTTAAA
GGATATACAACAACTCCACAAAAATGCTTGGAAGATTTTAAATTAGTCGTTTCTACATCT
CAATATGAAGGTCAAGGGTTAAGTATGATAGAAGCAATGATTTCTAAAAGGCCTGTTGTT
GCCTTTGACATCAAATACGGACCAAGTGATTTTATAGAAGATAATAAAAATGGTTATTTA
ATAGAAAACCATAATATTAACGACATGGCTGATAAAATACTTCAGCTTGTTAATAATGAT
GTATTAGCAGCGGAGTTTGGTTCGAAAGCGAGAGAAAACATTATAGAAAAATATTCAACG
GAATCAATATTAGAAAAATGGTTAAATCTTTTCAATAGCTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1482
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00973
- symbol: SAOUHSC_00973
- description:
- length: 493
- theoretical pI: 9.34538
- theoretical MW: 57274.4
- GRAVY: -0.49858
⊟Function[edit | edit source]
- ⊞⊞TIGRFAM: Protein fate Protein modification and repair accessory Sec system glycosylation protein GtfA (TIGR02918; EC 2.4.1.-; HMM-score: 117)and 15 more
- TheSEED :
- Poly(glycerol-phosphate) alpha-glucosyltransferase (EC 2.4.1.52)
- ⊞⊞PFAM: GT-B (CL0113) Glycos_transf_1; Glycosyl transferases group 1 (PF00534; HMM-score: 120.3)Glyco_trans_1_4; Glycosyl transferases group 1 (PF13692; HMM-score: 106)and 4 more
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKKIFMMVHELDVNKGGMTSSMFNRSKEFYDADIPADIVTFDYKGNYDEIIKALKKQGKMDRRTKMYNVFEYFKQISNNKHFKSNKLLYKHISERLKNTIEIEESKGISRYFDITTGTYIAYIRKSKSEKVIDFFKDNKRIERFSFIDNKVHMKETFNVDNKVCYQVFYDEKGYPYISRNINANNGAVGKTYVLVNKKEFKNNLALCVYYLEKLIKDSKDSIMICDGPGSFPKMFNTNHKNAQKYGVIHVNHHENFDDTGAFKKSEKYIIENANKINGVIVLTEAQRLDILNQFDVENIFTISNFVKIHNAPKHFQTEKIVGHISRMVPTKRIDLLIEVAELVVKKDNAVKFHIYGEGSVKDKIAKMIEDKNLERNVFLKGYTTTPQKCLEDFKLVVSTSQYEGQGLSMIEAMISKRPVVAFDIKYGPSDFIEDNKNGYLIENHNINDMADKILQLVNNDVLAAEFGSKARENIIEKYSTESILEKWLNLFNS
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [2] [3]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [4] : SAOUHSC_00972 > SAOUHSC_00973
⊟Regulation[edit | edit source]
- regulator:
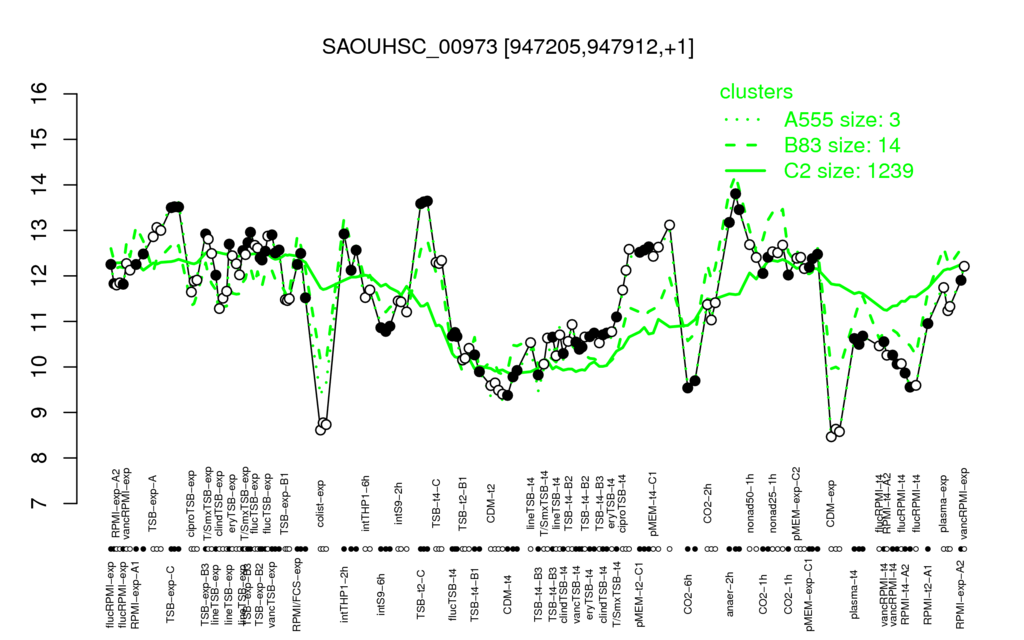
⊟Transcription pattern[edit | edit source]
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊞Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊞Other Information[edit | edit source]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Anne Berscheid, Peter Sass, Konstantin Weber-Lassalle, Ambrose L Cheung, Gabriele Bierbaum
Revisiting the genomes of the Staphylococcus aureus strains NCTC 8325 and RN4220.
Int J Med Microbiol: 2012, 302(2);84-7
[PubMed:22417616] [WorldCat.org] [DOI] (I p) - ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ Jump up to: 4.0 4.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
⊟Relevant publications[edit | edit source]
Solmaz Sobhanifar, Liam James Worrall, Robert J Gruninger, Gregory A Wasney, Markus Blaukopf, Lars Baumann, Emilie Lameignere, Matthew Solomonson, Eric D Brown, Stephen G Withers, Natalie C J Strynadka
Structure and mechanism of Staphylococcus aureus TarM, the wall teichoic acid α-glycosyltransferase.
Proc Natl Acad Sci U S A: 2015, 112(6);E576-85
[PubMed:25624472] [WorldCat.org] [DOI] (I p)