Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL2563 [new locus tag: SACOL_RS13425 ]
- pan locus tag?: SAUPAN006210000
- symbol: SACOL2563
- pan gene symbol?: clpL
- synonym:
- product: ATP-dependent Clp protease
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL2563 [new locus tag: SACOL_RS13425 ]
- symbol: SACOL2563
- product: ATP-dependent Clp protease
- replicon: chromosome
- strand: +
- coordinates: 2621890..2623995
- length: 2106
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238674 NCBI
- RefSeq: YP_187355 NCBI
- BioCyc: see SACOL_RS13425
- MicrobesOnline: 914034 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861
1921
1981
2041
2101ATGAATAACGGTTTTTTCAATAGCGACTTTGATTCAATTTTTCGAAGAATGATGAAAGAT
ATGCAAGGTTCAAATCAAGTCGGAAACAAAAAGTACTATATTAATGGTAAAGAAGTTTCA
CCTGAAGAACTAGCGCAACTCACACAACAAGGTGGCAATCACTCTGCTGAACAAAGTGCG
CAAGCTTTTCAACAAGCAGCACAAAGACAACAAGGGCAACAAGGTGGCAACGGCAATTAT
TTAGAACAAATTGGTCGTAACCTTACGCAAGAAGCACGTGACGGTTTATTAGATCCAGTC
ATTGGTCGTGATAAAGAAATTCAAGAAACTGCTGAAGTTTTAAGTAGACGAACTAAAAAC
AATCCTATATTAGTTGGAGAAGCTGGTGTTGGTAAAACTGCGATTGTTGAAGGTTTAGCA
CAGGCAATCGTTGAAGGAAATGTACCAGCAGCAATCAAAGACAAAGAAATTATTTCTGTA
GACATTTCATCATTAGAAGCTGGAACGCAATATCGTGGTGCTTTTGAAGAAAATATTCAA
AAATTAATCGAAGGTGTTAAATCTTCACAAAATGCCGTACTATTCTTTGATGAAATCCAT
CAAATTATCGGTTCAGGTGCCACAGGAAGTGATTCAGGTAGCAAAGGGTTATCTGATATT
TTGAAACCTGCATTAAGTCGTGGTGAGATTTCTATTATTGGTGCAACAACACAAGATGAA
TATCGAAACAATATTCTTAAAGATGCTGCATTAACGCGCAGATTTAATGAAGTGCTTGTT
AATGAACCAAGCGCTAAAGATACTGTTGAAATTTTAAAAGGTATTCGCGAAAAATTCGAA
GAACACCATCAAGTAAAATTACCAGATGACGTATTAAAAGCATGTGTTGACTTATCAATT
CAATATATTCCACAACGATTATTACCAGATAAAGCAATCGATGTGTTAGATATTACAGCA
GCACATTTATCTGCGCAAAGTCCAGCTGTCGATAAAGTTGAAACTGAAAAACGAATTTCT
GAATTAGAAAATGATAAACGTAAAGCAGTAAGTGCTGAAGAATATAAAAAAGCTGACGAC
ATTCAAAATGAAATCAAATCATTACAAGATAAATTAGAAAATAGTAATGGTGAACATACT
GCTGTTGCTACAGTTCATGATATTTCAGATACTATTCAACGATTAACTGGTATTCCAGTT
TCTCAAATGGATGATAACGATATTGAACGTTTAAAAAATATTTCTAATCGTTTAAGAAGT
AAAATCATAGGTCAAGATCAAGCTGTAGAAATGGTTTCACGTGCAATTCGCCGTAATCGT
GCTGGGTTTGATGACGGCAACCGTCCAATTGGCAGTTTCCTATTTGTTGGCCCTACTGGT
GTTGGTAAAACAGAGCTTGCTAAACAATTAGCAATTGATTTATTTGGTAATAAAGATGCA
CTGATTCGACTTGATATGAGTGAATATAGTGACACAACAGCTGTTTCAAAAATGATTGGT
ACAACTGCTGGTTATGTTGGTTATGATGACAATTCAAATACGTTAACTGAAAAAGTACGC
CGTAATCCATACTCAGTCATTCTATTTGATGAAATCGAAAAAGCAAATCCACAAATTTTA
ACATTGTTATTACAAGTAATGGATGATGGTAATTTGACTGATGGTCAAGGTAATGTCATC
AACTTTAAAAATACAATTATTATTTGTACATCAAATGCTGGCTTTGGCAATGGCAATGAC
GCTGAAGAAAAAGATATTATGCACGAAATGAAAAAATTCTTCCGCCCTGAATTCCTTAAC
CGCTTCAACGGCATCGTTGAATTCTTACATTTAGATAAAGATGCATTGCAAGATATCGTC
AACTTATTATTAGACGATGTACAAGTTACATTAGACAAAAAAGGTATTACGATGGACGTT
TCTCAAGATGCGAAAGATTGGTTAATTGAAGAAGGCTATGATGAAGAATTAGGTGCACGT
CCATTAAGACGTATTGTTGAACAGCAAGTACGTGACAAAATTACAGATTACTATTTAGAT
CATACAGACGTTAAACATGTGGATATAGATGTTGAGGATAACGAATTAGTCGTAAAAGGT
AAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1920
1980
2040
2100
2106
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL2563 [new locus tag: SACOL_RS13425 ]
- symbol: SACOL2563
- description: ATP-dependent Clp protease
- length: 701
- theoretical pI: 4.62942
- theoretical MW: 77835.7
- GRAVY: -0.506277
⊟Function[edit | edit source]
- ⊞TIGRFAM: Protein fate Protein folding and stabilization ATP-dependent chaperone protein ClpB (TIGR03346; HMM-score: 698)Protein fate Degradation of proteins, peptides, and glycopeptides ATP-dependent Clp protease ATP-binding subunit ClpA (TIGR02639; HMM-score: 668.9)Cellular processes Pathogenesis type VI secretion ATPase, ClpV1 family (TIGR03345; HMM-score: 603.2)Protein fate Protein and peptide secretion and trafficking type VI secretion ATPase, ClpV1 family (TIGR03345; HMM-score: 603.2)and 17 more
- TheSEED :
- ATP-dependent protease ATP-binding subunit clpL
- ⊞PFAM: P-loop_NTPase (CL0023) AAA_2; AAA domain (Cdc48 subfamily) (PF07724; HMM-score: 204.4)and 55 more
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MNNGFFNSDFDSIFRRMMKDMQGSNQVGNKKYYINGKEVSPEELAQLTQQGGNHSAEQSAQAFQQAAQRQQGQQGGNGNYLEQIGRNLTQEARDGLLDPVIGRDKEIQETAEVLSRRTKNNPILVGEAGVGKTAIVEGLAQAIVEGNVPAAIKDKEIISVDISSLEAGTQYRGAFEENIQKLIEGVKSSQNAVLFFDEIHQIIGSGATGSDSGSKGLSDILKPALSRGEISIIGATTQDEYRNNILKDAALTRRFNEVLVNEPSAKDTVEILKGIREKFEEHHQVKLPDDVLKACVDLSIQYIPQRLLPDKAIDVLDITAAHLSAQSPAVDKVETEKRISELENDKRKAVSAEEYKKADDIQNEIKSLQDKLENSNGEHTAVATVHDISDTIQRLTGIPVSQMDDNDIERLKNISNRLRSKIIGQDQAVEMVSRAIRRNRAGFDDGNRPIGSFLFVGPTGVGKTELAKQLAIDLFGNKDALIRLDMSEYSDTTAVSKMIGTTAGYVGYDDNSNTLTEKVRRNPYSVILFDEIEKANPQILTLLLQVMDDGNLTDGQGNVINFKNTIIICTSNAGFGNGNDAEEKDIMHEMKKFFRPEFLNRFNGIVEFLHLDKDALQDIVNLLLDDVQVTLDKKGITMDVSQDAKDWLIEEGYDEELGARPLRRIVEQQVRDKITDYYLDHTDVKHVDIDVEDNELVVKGK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3]
- quantitative data / protein copy number per cell: 1136 [4]
- ⊟interaction partners:
SACOL1800 (dat) D-alanine aminotransferase [5] (data from MRSA252) SACOL1329 (femC) glutamine synthetase [5] (data from MRSA252) SACOL1734 (gapA2) glyceraldehyde 3-phosphate dehydrogenase 2 [5] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [5] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [5] (data from MRSA252) SACOL2233 (rpsC) 30S ribosomal protein S3 [5] (data from MRSA252) SACOL2222 (rpsE) 30S ribosomal protein S5 [5] (data from MRSA252) SACOL0944 NADH dehydrogenase [5] (data from MRSA252) SACOL1759 universal stress protein [5] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
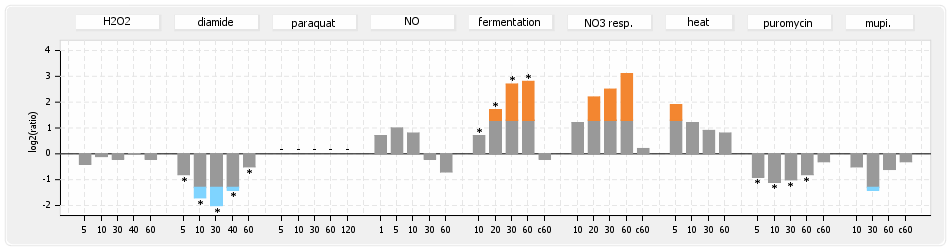
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊞Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊞Other Information[edit | edit source]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Jump up to: 5.0 5.1 5.2 5.3 5.4 5.5 5.6 5.7 5.8 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Markus Bischoff, Paul Dunman, Jan Kormanec, Daphne Macapagal, Ellen Murphy, William Mounts, Brigitte Berger-Bächi, Steven Projan
Microarray-based analysis of the Staphylococcus aureus sigmaB regulon.
J Bacteriol: 2004, 186(13);4085-99
[PubMed:15205410] [WorldCat.org] [DOI] (P p) - ↑ Jan Pané-Farré, Beate Jonas, Konrad Förstner, Susanne Engelmann, Michael Hecker
The sigmaB regulon in Staphylococcus aureus and its regulation.
Int J Med Microbiol: 2006, 296(4-5);237-58
[PubMed:16644280] [WorldCat.org] [DOI] (P p)