Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02244
- pan locus tag?: SAUPAN005193000
- symbol: SAOUHSC_02244
- pan gene symbol?: —
- synonym:
- product: succinyl-diaminopimelate desuccinylase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02244
- symbol: SAOUHSC_02244
- product: succinyl-diaminopimelate desuccinylase
- replicon: chromosome
- strand: +
- coordinates: 2078221..2079444
- length: 1224
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919665 NCBI
- RefSeq: YP_500727 NCBI
- BioCyc: G1I0R-2118 BioCyc
- MicrobesOnline: 1290683 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201ATGACAACTTTTAGTGAAAAAGAAAAAATTCAATTACTAGCAGATATTGTTGAACTACAA
ACTGAAAATAATAATGAAATAGACGTTTGTAATTATTTAACAGATTTATTCGACAAGTAC
GATATTAAATCTGAAATTTTGAAAGTTAATGAACACCGCGCCAATATCGTTGCAGAAATC
GGTAACGGCTCACCTATACTCGCATTGAGTGGTCATATGGATGTTGTTGATGCAGGAAAT
CAAGATAATTGGTCATATCCCCCTTTTCAACTGACAGAAAAAGATGGCAAATTATATGGC
CGAGGCACTACAGATATGAAAGGCGGTTTAATGGCTTTGGTCGTATCTCTAATCGAATTA
AAAGAACAAAATGAATTGCCTCATGGAACGATTAGATTACTGGCTACTGCTGGCGAAGAG
AAAGAACAAGAAGGTGCCAAATTATTAGCTGATAAAGGCTATTTAGACGATGTCGATGGC
TTAATTATTGCTGAACCAACTGGATCTGGAATTTATTATGCACATAAGGGGTCTATGTCA
TGTAAAGTAACTGCAACTGGTAAAGCTGTCCATAGCTCAGTTCCATTTATTGGTGACAAT
GCAATTGATACACTGCTTGAATTTTATAATCTATTTAAAGAAAAATATTCAGAGCTTAAA
CAACAAGATACTAAACATGAATTAGATGTTGCGCCTATGTTCAAATCATTGATTGGAAAA
GAAATTTCTGAAGAGGATGCAAATTATGCATCTGGTCTTACAGCTGTATGTTCGATTATA
AATGGCGGCAAACAATTTAACTCTGTACCAGATGAAGCTTCACTTGAATTTAACGTAAGA
CCAGTTCCTGAGTATGATAACGACTTTATAGAATCGTTTTTCCAAAATATCATTAATGAT
GTGGATAGCAATAAGCTTTCACTCGATATTCCAAGCAATCACCGACCTGTAACAAGCGAT
AAAAATAGCAAATTAATTACTACGATTAAAGATGTAGCTTCTAGTTATGTAGAACAAGAC
GAAATATTTGTTTCAGCGCTTGTAGGCGCAACAGATGCCTCTAGTTTCTTAGGAGATAAT
AAGGACAATGTTGATTTAGCCATTTTTGGACCAGGTAATCCATTAATGGCACATCAAATC
GATGAATATATTGAAAAAGATATGTATCTGAAATATATTGATATTTTTAAAGAGGCTTCC
ATTCAATATTTAAAAGAAAAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1224
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02244
- symbol: SAOUHSC_02244
- description: succinyl-diaminopimelate desuccinylase
- length: 407
- theoretical pI: 4.27856
- theoretical MW: 45097.3
- GRAVY: -0.329975
⊟Function[edit | edit source]
- TIGRFAM: Protein fate Degradation of proteins, peptides, and glycopeptides peptidase, ArgE/DapE family (TIGR01910; EC 3.4.-.-; HMM-score: 300.7)and 11 moreAmino acid biosynthesis Glutamate family acetylornithine deacetylase (ArgE) (TIGR01892; EC 3.5.1.16; HMM-score: 167.9)Amino acid biosynthesis Aspartate family succinyl-diaminopimelate desuccinylase (TIGR01246; EC 3.5.1.18; HMM-score: 134.5)Amino acid biosynthesis Aspartate family succinyl-diaminopimelate desuccinylase (TIGR01900; EC 3.5.1.18; HMM-score: 122)putative selenium metabolism hydrolase (TIGR03526; HMM-score: 114.6)N-acetyl-ornithine/N-acetyl-lysine deacetylase (TIGR01902; HMM-score: 100.7)putative dipeptidase (TIGR01887; EC 3.4.13.-; HMM-score: 95.3)N-acyl-L-amino-acid amidohydrolase (TIGR01880; EC 3.5.1.14; HMM-score: 78.3)Protein fate Degradation of proteins, peptides, and glycopeptides amidohydrolase (TIGR01891; HMM-score: 75.6)Protein fate Degradation of proteins, peptides, and glycopeptides dipeptidase PepV (TIGR01886; EC 3.4.13.-; HMM-score: 51.8)peptidase T-like protein (TIGR01883; HMM-score: 31.3)Xaa-His dipeptidase (TIGR01893; EC 3.4.13.20; HMM-score: 29.2)
- TheSEED :
- Acetylornithine deacetylase (EC 3.5.1.16)
- PFAM: ZnExoPePases (CL0865) Peptidase_M20; Peptidase family M20/M25/M40 (PF01546; HMM-score: 133.6)and 2 moreno clan defined M20_dimer; Peptidase dimerisation domain (PF07687; HMM-score: 78.6)ZnExoPePases (CL0865) Peptidase_M28; Peptidase family M28 (PF04389; HMM-score: 27.1)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9635
- Cytoplasmic Membrane Score: 0.0242
- Cell wall & surface Score: 0.0025
- Extracellular Score: 0.0097
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.003302
- TAT(Tat/SPI): 0.000133
- LIPO(Sec/SPII): 0.000511
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTTFSEKEKIQLLADIVELQTENNNEIDVCNYLTDLFDKYDIKSEILKVNEHRANIVAEIGNGSPILALSGHMDVVDAGNQDNWSYPPFQLTEKDGKLYGRGTTDMKGGLMALVVSLIELKEQNELPHGTIRLLATAGEEKEQEGAKLLADKGYLDDVDGLIIAEPTGSGIYYAHKGSMSCKVTATGKAVHSSVPFIGDNAIDTLLEFYNLFKEKYSELKQQDTKHELDVAPMFKSLIGKEISEEDANYASGLTAVCSIINGGKQFNSVPDEASLEFNVRPVPEYDNDFIESFFQNIINDVDSNKLSLDIPSNHRPVTSDKNSKLITTIKDVASSYVEQDEIFVSALVGATDASSFLGDNKDNVDLAIFGPGNPLMAHQIDEYIEKDMYLKYIDIFKEASIQYLKEK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization:
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator: SigB* (activation) regulon
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
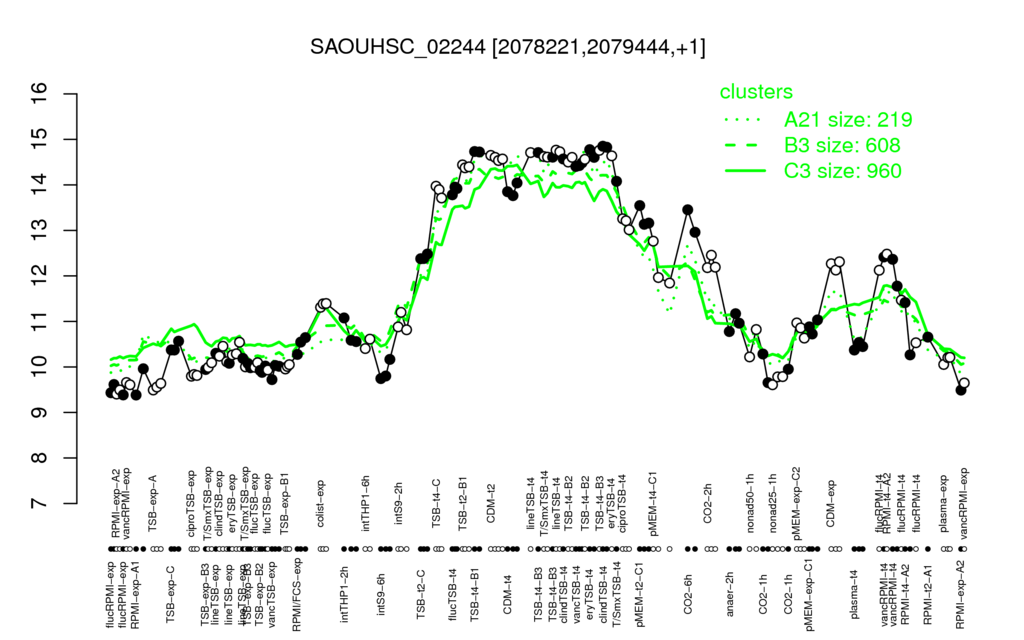 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ Markus Bischoff, Paul Dunman, Jan Kormanec, Daphne Macapagal, Ellen Murphy, William Mounts, Brigitte Berger-Bächi, Steven Projan
Microarray-based analysis of the Staphylococcus aureus sigmaB regulon.
J Bacteriol: 2004, 186(13);4085-99
[PubMed:15205410] [WorldCat.org] [DOI] (P p) - ↑ 4.0 4.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
