Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL0586 [new locus tag: SACOL_RS03040 ]
- pan locus tag?: SAUPAN002311000
- symbol: rplL
- pan gene symbol?: rplL
- synonym:
- product: 50S ribosomal protein L7/L12
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL0586 [new locus tag: SACOL_RS03040 ]
- symbol: rplL
- product: 50S ribosomal protein L7/L12
- replicon: chromosome
- strand: +
- coordinates: 605622..605990
- length: 369
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3236232 NCBI
- RefSeq: YP_185472 NCBI
- BioCyc: see SACOL_RS03040
- MicrobesOnline: 912066 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361ATGGCTAATCATGAACAAATCATTGAAGCGATTAAAGAAATGTCAGTATTAGAATTAAAC
GACTTAGTAAAAGCAATTGAAGAAGAATTTGGTGTAACTGCAGCTGCTCCAGTAGCAGTA
GCAGGTGCAGCTGGTGGCGCTGACGCTGCAGCAGAAAAAACTGAATTTGACGTTGAGTTA
ACTTCAGCTGGTTCATCTAAAATCAAAGTTGTTAAAGCTGTTAAAGAAGCAACTGGTTTA
GGATTAAAAGATGCTAAAGAATTAGTAGACGGAGCTCCTAAAGTAATCAAAGAAGCTTTA
CCTAAAGAAGAAGCTGAAAAACTTAAAGAACAATTAGAAGAAGTTGGAGCTACTGTAGAA
TTAAAATAA60
120
180
240
300
360
369
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL0586 [new locus tag: SACOL_RS03040 ]
- symbol: RplL
- description: 50S ribosomal protein L7/L12
- length: 122
- theoretical pI: 4.32374
- theoretical MW: 12711.5
- GRAVY: -0.0483607
⊟Function[edit | edit source]
- TIGRFAM: Protein synthesis Ribosomal proteins: synthesis and modification ribosomal protein bL12 (TIGR00855; HMM-score: 158.6)
- TheSEED :
- LSU ribosomal protein L7p/L12p (P1/P2)
- PFAM: no clan defined Ribosomal_L12; Ribosomal protein L7/L12 C-terminal domain (PF00542; HMM-score: 104.3)and 6 moreRibosomal_L12_N; Ribosomal protein L7/L12 dimerisation domain (PF16320; HMM-score: 75.5)DUF2267; Uncharacterized conserved protein (DUF2267) (PF10025; HMM-score: 16.3)IHF-likeDNA-bdg (CL0548) Bac_DNA_binding; Bacterial DNA-binding protein (PF00216; HMM-score: 15.7)no clan defined Ribosomal_60s; 60s Acidic ribosomal protein (PF00428; HMM-score: 15)CwsA; Cell wall synthesis protein CwsA (PF10814; HMM-score: 14.5)HTH (CL0123) HTH_33; Winged helix-turn helix (PF13592; HMM-score: 12.3)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9475
- Cytoplasmic Membrane Score: 0.0177
- Cell wall & surface Score: 0.0002
- Extracellular Score: 0.0346
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 0.67
- Signal peptide possibility: -0.5
- N-terminally Anchored Score: -2
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.057639
- TAT(Tat/SPI): 0.076736
- LIPO(Sec/SPII): 0.001562
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MANHEQIIEAIKEMSVLELNDLVKAIEEEFGVTAAAPVAVAGAAGGADAAAEKTEFDVELTSAGSSKIKVVKAVKEATGLGLKDAKELVDGAPKVIKEALPKEEAEKLKEQLEEVGATVELK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3] [4] [5]
- quantitative data / protein copy number per cell: 8346 [6]
- interaction partners:
SACOL1760 (ackA) acetate kinase [7] (data from MRSA252) SACOL0452 (ahpC) alkyl hydroperoxide reductase subunit C [7] (data from MRSA252) SACOL0557 (cysK) cysteine synthase [7] (data from MRSA252) SACOL1637 (dnaK) molecular chaperone DnaK [7] (data from MRSA252) SACOL0842 (eno) phosphopyruvate hydratase [7] (data from MRSA252) SACOL2622 (fdaB) fructose-1,6-bisphosphate aldolase [7] (data from MRSA252) SACOL1199 (ftsZ) cell division protein FtsZ [7] (data from MRSA252) SACOL0593 (fusA) elongation factor G [7] (data from MRSA252) SACOL0838 (gapA1) glyceraldehyde 3-phosphate dehydrogenase [7] (data from MRSA252) SACOL0460 (guaB) inosine-5'-monophosphate dehydrogenase [7] (data from MRSA252) SACOL1513 (hup) DNA-binding protein HU [7] (data from MRSA252) SACOL1741 (icd) isocitrate dehydrogenase [7] (data from MRSA252) SACOL2618 (ldh2) L-lactate dehydrogenase [7] (data from MRSA252) SACOL2623 (mqo2) malate:quinone oxidoreductase [7] (data from MRSA252) SACOL0746 (norR) MarR family transcriptional regulator [7] (data from MRSA252) SACOL1102 (pdhA) pyruvate dehydrogenase complex E1 component subunit alpha [7] (data from MRSA252) SACOL1103 (pdhB) pyruvate dehydrogenase complex E1 component subunit beta [7] (data from MRSA252) SACOL1104 (pdhC) branched-chain alpha-keto acid dehydrogenase E2 [7] (data from MRSA252) SACOL1105 (pdhD) dihydrolipoamide dehydrogenase [7] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [7] (data from MRSA252) SACOL0204 (pflB) formate acetyltransferase [7] (data from MRSA252) SACOL0839 (pgk) phosphoglycerate kinase [7] (data from MRSA252) SACOL1293 (pnp) polynucleotide phosphorylase/polyadenylase [7] (data from MRSA252) SACOL1092 (ptsI) phosphoenolpyruvate-protein phosphotransferase [7] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [7] (data from MRSA252) SACOL0584 (rplA) 50S ribosomal protein L1 [7] (data from MRSA252) SACOL2236 (rplB) 50S ribosomal protein L2 [7] (data from MRSA252) SACOL2239 (rplC) 50S ribosomal protein L3 [7] (data from MRSA252) SACOL2238 (rplD) 50S ribosomal protein L4 [7] (data from MRSA252) SACOL2224 (rplF) 50S ribosomal protein L6 [7] (data from MRSA252) SACOL0585 (rplJ) 50S ribosomal protein L10 [7] (data from MRSA252) SACOL0583 (rplK) 50S ribosomal protein L11 [7] (data from MRSA252) SACOL2207 (rplM) 50S ribosomal protein L13 [7] (data from MRSA252) SACOL2220 (rplO) 50S ribosomal protein L15 [7] (data from MRSA252) SACOL2223 (rplR) 50S ribosomal protein L18 [7] (data from MRSA252) SACOL1257 (rplS) 50S ribosomal protein L19 [7] (data from MRSA252) SACOL1725 (rplT) 50S ribosomal protein L20 [7] (data from MRSA252) SACOL1702 (rplU) 50S ribosomal protein L21 [7] (data from MRSA252) SACOL2234 (rplV) 50S ribosomal protein L22 [7] (data from MRSA252) SACOL2221 (rpmD) 50S ribosomal protein L30 [7] (data from MRSA252) SACOL0588 (rpoB) DNA-directed RNA polymerase subunit beta [7] (data from MRSA252) SACOL1516 (rpsA) 30S ribosomal protein S1 [7] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [7] (data from MRSA252) SACOL2233 (rpsC) 30S ribosomal protein S3 [7] (data from MRSA252) SACOL2222 (rpsE) 30S ribosomal protein S5 [7] (data from MRSA252) SACOL0437 (rpsF) 30S ribosomal protein S6 [7] (data from MRSA252) SACOL0592 (rpsG) 30S ribosomal protein S7 [7] (data from MRSA252) SACOL2206 (rpsI) 30S ribosomal protein S9 [7] (data from MRSA252) SACOL2214 (rpsK) 30S ribosomal protein S11 [7] (data from MRSA252) SACOL0591 (rpsL) 30S ribosomal protein S12 [7] (data from MRSA252) SACOL2215 (rpsM) 30S ribosomal protein S13 [7] (data from MRSA252) SACOL0816 (secA) preprotein translocase subunit SecA [7] (data from MRSA252) SACOL1610 (sodA2) superoxide dismutase [7] (data from MRSA252) SACOL0095 (spa) immunoglobulin G binding protein A precursor [7] (data from MRSA252) SACOL1449 (sucA) 2-oxoglutarate dehydrogenase E1 component [7] (data from MRSA252) SACOL1448 (sucB) dihydrolipoamide succinyltransferase [7] (data from MRSA252) SACOL1831 (tal) translaldolase [7] (data from MRSA252) SACOL1377 (tkt) transketolase [7] (data from MRSA252) SACOL0840 (tpiA) triosephosphate isomerase [7] (data from MRSA252) SACOL1276 (tsf) elongation factor Ts [7] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [7] (data from MRSA252) SACOL0303 5'-nucleotidase [7] (data from MRSA252) SACOL0944 NADH dehydrogenase [7] (data from MRSA252) SACOL0973 fumarylacetoacetate hydrolase [7] (data from MRSA252) SACOL1437 CSD family cold shock protein [7] (data from MRSA252) SACOL1753 universal stress protein [7] (data from MRSA252) SACOL1759 universal stress protein [7] (data from MRSA252) SACOL1836 hypothetical protein [7] (data from MRSA252) SACOL1952 ferritins family protein [7] (data from MRSA252) SACOL1973 hypothetical protein [7] (data from MRSA252) SACOL2163 hypothetical protein [7] (data from MRSA252) SACOL2173 alkaline shock protein 23 [7] (data from MRSA252) SACOL2266 molybdopterin biosynthesis MoeA protein [7] (data from MRSA252) SACOL2469 glutamate synthase [7] (data from MRSA252) SACOL2569 1-pyrroline-5-carboxylate dehydrogenase [7] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: L10 leader (transcription termination) regulon
L10 leader (RNA) important in Ribosome biogenesis; transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
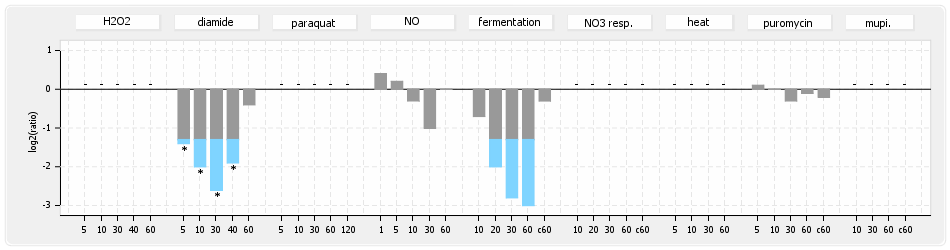
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 19.63 h [8]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Annette Dreisbach, Kristina Hempel, Girbe Buist, Michael Hecker, Dörte Becher, Jan Maarten van Dijl
Profiling the surfacome of Staphylococcus aureus.
Proteomics: 2010, 10(17);3082-96
[PubMed:20662103] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 7.00 7.01 7.02 7.03 7.04 7.05 7.06 7.07 7.08 7.09 7.10 7.11 7.12 7.13 7.14 7.15 7.16 7.17 7.18 7.19 7.20 7.21 7.22 7.23 7.24 7.25 7.26 7.27 7.28 7.29 7.30 7.31 7.32 7.33 7.34 7.35 7.36 7.37 7.38 7.39 7.40 7.41 7.42 7.43 7.44 7.45 7.46 7.47 7.48 7.49 7.50 7.51 7.52 7.53 7.54 7.55 7.56 7.57 7.58 7.59 7.60 7.61 7.62 7.63 7.64 7.65 7.66 7.67 7.68 7.69 7.70 7.71 7.72 7.73 7.74 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)