m (Text replacement - "gene Genbank" to "gene RefSeq") |
m (Text replacement - "* <aureodatabase>protein Genbank</aureodatabase> " to "") |
||
| Line 1: | Line 1: | ||
__TOC__ | |||
<protect> | <protect> | ||
<aureodatabase> | <aureodatabase>annotation</aureodatabase> | ||
=Summary= | =Summary= | ||
* <aureodatabase>organism</aureodatabase> | *<aureodatabase>organism</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>pan locus</aureodatabase> | *<aureodatabase>pan locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>pan gene symbol</aureodatabase> | *<aureodatabase>pan gene symbol</aureodatabase> | ||
* <aureodatabase>gene synonyms</aureodatabase> | *<aureodatabase>gene synonyms</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
</protect> | </protect> | ||
| Line 24: | Line 25: | ||
==General== | ==General== | ||
* <aureodatabase>gene type</aureodatabase> | *<aureodatabase>gene type</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
* <aureodatabase>gene replicon</aureodatabase> | *<aureodatabase>gene replicon</aureodatabase> | ||
* <aureodatabase>strand</aureodatabase> | *<aureodatabase>strand</aureodatabase> | ||
* <aureodatabase>gene coordinates</aureodatabase> | *<aureodatabase>gene coordinates</aureodatabase> | ||
* <aureodatabase>gene length</aureodatabase> | *<aureodatabase>gene length</aureodatabase> | ||
* <aureodatabase>essential</aureodatabase> | *<aureodatabase>essential</aureodatabase> | ||
*<aureodatabase>gene comment</aureodatabase> | |||
</protect> | </protect> | ||
| Line 38: | Line 40: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>gene GI</aureodatabase> | *<aureodatabase>gene GI</aureodatabase> | ||
* <aureodatabase>gene RefSeq</aureodatabase> | *<aureodatabase>gene RefSeq</aureodatabase> | ||
*<aureodatabase>gene BioCyc</aureodatabase> | |||
*<aureodatabase>gene MicrobesOnline</aureodatabase> | |||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Phenotype== | ==Phenotype== | ||
</protect> | </protect> | ||
Share your knowledge and add information here. [<span class="plainlinks">[//aureowiki.med.uni-greifswald.de/index.php?title={{PAGENAMEE}}&veaction=edit§ion=6 edit]</span>] | |||
<protect> | <protect> | ||
==DNA sequence== | ==DNA sequence== | ||
* <aureodatabase>gene sequence</aureodatabase> | *<aureodatabase>gene sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
<aureodatabase>RNA regulated operons</aureodatabase> | |||
</protect> | |||
<protect> | |||
=Protein= | =Protein= | ||
<aureodatabase>protein 3D view</aureodatabase> | <aureodatabase>protein 3D view</aureodatabase> | ||
==General== | ==General== | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>protein symbol</aureodatabase> | *<aureodatabase>protein symbol</aureodatabase> | ||
* <aureodatabase>protein description</aureodatabase> | *<aureodatabase>protein description</aureodatabase> | ||
* <aureodatabase>protein length</aureodatabase> | *<aureodatabase>protein length</aureodatabase> | ||
* <aureodatabase>theoretical pI</aureodatabase> | *<aureodatabase>theoretical pI</aureodatabase> | ||
* <aureodatabase>theoretical MW</aureodatabase> | *<aureodatabase>theoretical MW</aureodatabase> | ||
* <aureodatabase>GRAVY</aureodatabase> | *<aureodatabase>GRAVY</aureodatabase> | ||
</protect> | </protect> | ||
| Line 71: | Line 78: | ||
==Function== | ==Function== | ||
* <aureodatabase>protein reaction</aureodatabase> | *<aureodatabase>protein reaction</aureodatabase> | ||
* <aureodatabase>protein TIGRFAM</aureodatabase> | *<aureodatabase>protein TIGRFAM</aureodatabase> | ||
* <aureodatabase>protein TheSeed</aureodatabase> | *<aureodatabase>protein TheSeed</aureodatabase> | ||
* <aureodatabase>protein PFAM</aureodatabase> | *<aureodatabase>protein PFAM</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Structure, modifications & | ==Structure, modifications & cofactors== | ||
* <aureodatabase>protein domains</aureodatabase> | *<aureodatabase>protein domains</aureodatabase> | ||
* <aureodatabase>protein modifications</aureodatabase> | *<aureodatabase>protein modifications</aureodatabase> | ||
* <aureodatabase>protein cofactors</aureodatabase> | *<aureodatabase>protein cofactors</aureodatabase> | ||
* <aureodatabase>protein effectors</aureodatabase> | *<aureodatabase>protein effectors</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein regulated operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 90: | Line 97: | ||
==Localization== | ==Localization== | ||
* <aureodatabase>protein Psortb</aureodatabase> | *<aureodatabase>protein Psortb</aureodatabase> | ||
* <aureodatabase>protein LocateP</aureodatabase> | *<aureodatabase>protein LocateP</aureodatabase> | ||
* <aureodatabase>protein SignalP</aureodatabase> | *<aureodatabase>protein SignalP</aureodatabase> | ||
* <aureodatabase>protein TMHMM</aureodatabase> | *<aureodatabase>protein TMHMM</aureodatabase> | ||
</protect> | </protect> | ||
| Line 99: | Line 106: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>protein GI</aureodatabase> | *<aureodatabase>protein GI</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein RefSeq</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein UniProt</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein STRING</aureodatabase> | ||
</protect> | </protect> | ||
| Line 108: | Line 115: | ||
==Protein sequence== | ==Protein sequence== | ||
* <aureodatabase>protein sequence</aureodatabase> | *<aureodatabase>protein sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Experimental data== | ||
* <aureodatabase>protein validated peptides</aureodatabase> | *<aureodatabase>protein validated peptides</aureodatabase> | ||
*<aureodatabase>protein validated localization</aureodatabase> | |||
*<aureodatabase>protein validated quantitative data</aureodatabase> | |||
*<aureodatabase>protein partners</aureodatabase> | |||
</protect> | </protect> | ||
| Line 125: | Line 135: | ||
==Operon== | ==Operon== | ||
* <aureodatabase>operons</aureodatabase> | *<aureodatabase>operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 131: | Line 141: | ||
==Regulation== | ==Regulation== | ||
*<aureodatabase>regulators</aureodatabase> | |||
* <aureodatabase>regulators</aureodatabase> | |||
</protect> | </protect> | ||
| Line 138: | Line 147: | ||
==Transcription pattern== | ==Transcription pattern== | ||
* <aureodatabase>expression browser</aureodatabase> | *<aureodatabase>expression browser</aureodatabase> | ||
</protect> | </protect> | ||
| Line 144: | Line 153: | ||
==Protein synthesis (provided by Aureolib)== | ==Protein synthesis (provided by Aureolib)== | ||
* <aureodatabase>protein synthesis Aureolib</aureodatabase> | *<aureodatabase>protein synthesis Aureolib</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Protein stability== | ||
* <aureodatabase>protein half-life</aureodatabase> | *<aureodatabase>protein half-life</aureodatabase> | ||
</protect> | </protect> | ||
Latest revision as of 08:39, 11 March 2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_03002
- pan locus tag?: SAUPAN006411000
- symbol: SAOUHSC_03002
- pan gene symbol?: icaA
- synonym:
- product: N-glycosyltransferase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_03002
- symbol: SAOUHSC_03002
- product: N-glycosyltransferase
- replicon: chromosome
- strand: +
- coordinates: 2775174..2776412
- length: 1239
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921484 NCBI
- RefSeq: YP_501451 NCBI
- BioCyc: G1I0R-2824 BioCyc
- MicrobesOnline: 1291422 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201TTGCAATTTTTTAACTTTTTGCTTTTTTATCCTGTATTTATGTCTATTTACTGGATTGTC
GGTTCAATTTATTTCTATTTTACCAGAGAAATTAGATATTCATTGAACAAGAAGCCTGAC
ATAAATGTGGATGAATTAGAAGGCATTACATTTTTACTTGCCTGTTATAACGAAAGTGAA
ACGATTGAAGATACGTTGTCTAATGTTCTTGCACTCAAATACGAGAAGAAAGAAATTATT
ATCATTAATGATGGAAGTTCAGATAATACAGCAGAACTCATCTATAAAATCAAAGAAAAT
AATGACTTTATTTTCGTCGATTTACAAGAAAACAGAGGTAAAGCCAACGCACTCAATCAA
GGCATTAAACAGGCTTCATATGATTATGTAATGTGCTTGGATGCAGATACTATCGTTGAT
CAAGATGCACCATATTATATGATTGAGAATTTCAAACATGATCCAAAACTTGGTGCAGTT
ACAGGTAATCCTAGAATTCGAAATAAGAGTTCTATTTTAGGTAAAATTCAAACGATAGAA
TATGCAAGTTTAATTGGCTGTATTAAGCGAAGTCAGACACTTGCTGGCGCAGTCAATACT
ATTTCGGGTGTCTTCACTCTATTTAAAAAAAGTGCAGTTGTCGACGTTGGCTACTGGGAT
ACTGATATGATTACCGAAGATATTGCAGTTTCTTGGAAATTGCATTTACGTGGATATCGT
ATTAAGTATGAACCGCTTGCCATGTGTTGGATGTTGGTTCCAGAAACATTGGGAGGTCTT
TGGAAGCAACGCGTGAGATGGGCTCAAGGGGGACACGAAGTATTACTACGAGACTTTTTT
AGCACAATGAAAACGAAAAGGTTTCCTTTATATATTTTGATGTTTGAGCAAATCATCTCG
ATTTTATGGGTATATATAGTGCTTCTATATTTAGGCTATTTGTTCATAACAGCAAACTTC
TTAGACTATACATTTATGACATATAGTTTTTCAATATTTCTACTATCATCATTTACTATG
ACTTTTATAAACGTTATTCAATTTACAGTCGCACTCTTTATTGATAGTCGCTACGAGAAA
AAGAATATGGCTGGACTCATATTTGTAAGTTGGTATCCGACAGTATACTGGATTATTAAC
GCAGCAGTAGTTCTTGTCGCATTTCCAAAAGCATTAAAACGTAAGAAAGGTGGTTACGCA
ACATGGTCAAGCCCAGACAGAGGGAATACCCAACGCTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1239
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_03002
- symbol: SAOUHSC_03002
- description: N-glycosyltransferase
- length: 412
- theoretical pI: 8.45984
- theoretical MW: 47769.2
- GRAVY: 0.112864
⊟Function[edit | edit source]
- reaction: EC 2.4.1.-? ExPASy
- TIGRFAM: poly-beta-1,6 N-acetyl-D-glucosamine synthase (TIGR03937; EC 2.4.1.-; HMM-score: 633.2)and 11 moreputative glycosyltransferase, exosortase G-associated (TIGR03111; HMM-score: 98.7)chitooligosaccharide synthase NodC (TIGR04242; EC 2.4.1.-; HMM-score: 85.8)mycofactocin system glycosyltransferase (TIGR03965; HMM-score: 57.5)cellulose synthase catalytic subunit (UDP-forming) (TIGR03030; EC 2.4.1.12; HMM-score: 50.2)Unknown function Enzymes of unknown specificity transferase 2, rSAM/selenodomain-associated (TIGR04283; EC 2.-.-.-; HMM-score: 49.7)glycosyltransferase, TIGR04182 family (TIGR04182; HMM-score: 32.7)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides rhamnosyltransferase (TIGR01556; EC 2.4.1.-; HMM-score: 28.8)hopene-associated glycosyltransferase HpnB (TIGR03469; HMM-score: 28)colanic acid biosynthesis glycosyltransferase WcaE (TIGR04009; EC 2.4.-.-; HMM-score: 22.6)hopanoid biosynthesis associated glycosyl transferase protein HpnI (TIGR03472; HMM-score: 20.4)colanic acid biosynthesis glycosyltransferase WcaA (TIGR04017; EC 2.4.-.-; HMM-score: 17.7)
- TheSEED :
- Polysaccharide intercellular adhesin (PIA) biosynthesis N-glycosyltransferase IcaA (EC 2.4.-.-)
- PFAM: GT-A (CL0110) Glycos_transf_2; Glycosyl transferase family 2 (PF00535; HMM-score: 105.1)Glyco_trans_2_3; Glycosyl transferase family group 2 (PF13632; HMM-score: 93.6)Glyco_tranf_2_3; Glycosyltransferase like family 2 (PF13641; HMM-score: 92.3)and 8 moreGlyco_transf_21; Glycosyl transferase family 21 (PF13506; HMM-score: 68.6)Chitin_synth_2; Chitin synthase (PF03142; HMM-score: 49.6)Glyco_tranf_2_2; Glycosyltransferase like family 2 (PF10111; HMM-score: 39.4)Glyco_tranf_2_4; Glycosyl transferase family 2 (PF13704; HMM-score: 28)Glyco_tranf_2_5; Glycosyltransferase like family (PF13712; HMM-score: 24.2)Cellulose_synt; Cellulose synthase (PF03552; HMM-score: 22.5)AMP_N-like (CL0356) Creatinase_N_2; Creatinase/Prolidase N-terminal domain (PF16189; HMM-score: 17.9)no clan defined DUF393; Protein of unknown function, DUF393 (PF04134; HMM-score: 15.1)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 4
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.023665
- TAT(Tat/SPI): 0.000501
- LIPO(Sec/SPII): 0.029446
- predicted transmembrane helices (TMHMM): 4
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MQFFNFLLFYPVFMSIYWIVGSIYFYFTREIRYSLNKKPDINVDELEGITFLLACYNESETIEDTLSNVLALKYEKKEIIIINDGSSDNTAELIYKIKENNDFIFVDLQENRGKANALNQGIKQASYDYVMCLDADTIVDQDAPYYMIENFKHDPKLGAVTGNPRIRNKSSILGKIQTIEYASLIGCIKRSQTLAGAVNTISGVFTLFKKSAVVDVGYWDTDMITEDIAVSWKLHLRGYRIKYEPLAMCWMLVPETLGGLWKQRVRWAQGGHEVLLRDFFSTMKTKRFPLYILMFEQIISILWVYIVLLYLGYLFITANFLDYTFMTYSFSIFLLSSFTMTFINVIQFTVALFIDSRYEKKNMAGLIFVSWYPTVYWIINAAVVLVAFPKALKRKKGGYATWSSPDRGNTQR
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization:
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulators: CodY* (repression) regulon, IcaR* (repression) regulon
CodY* (TF) important in Amino acid metabolism; RegPrecise IcaR* (TF) important in Intercellular adhesion; RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
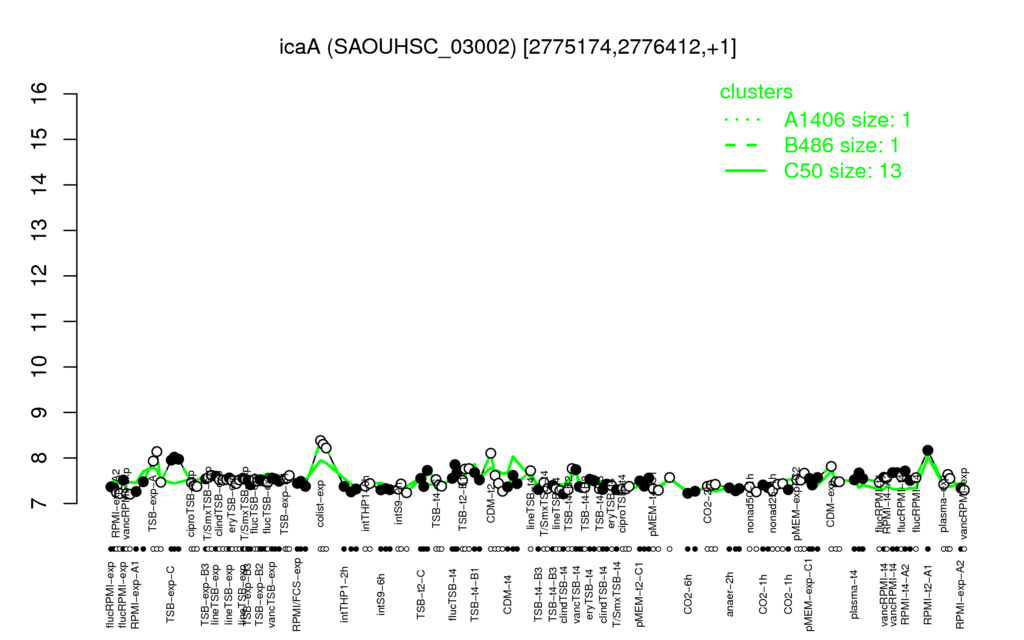 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
⊟Relevant publications[edit | edit source]
S E Cramton, C Gerke, N F Schnell, W W Nichols, F Götz
The intercellular adhesion (ica) locus is present in Staphylococcus aureus and is required for biofilm formation.
Infect Immun: 1999, 67(10);5427-33
[PubMed:10496925] [WorldCat.org] [DOI] (P p)Ursula Fluckiger, Martina Ulrich, Andrea Steinhuber, Gerd Döring, Dietrich Mack, Regine Landmann, Christiane Goerke, Christiane Wolz
Biofilm formation, icaADBC transcription, and polysaccharide intercellular adhesin synthesis by staphylococci in a device-related infection model.
Infect Immun: 2005, 73(3);1811-9
[PubMed:15731082] [WorldCat.org] [DOI] (P p)C Heilmann, O Schweitzer, C Gerke, N Vanittanakom, D Mack, F Götz
Molecular basis of intercellular adhesion in the biofilm-forming Staphylococcus epidermidis.
Mol Microbiol: 1996, 20(5);1083-91
[PubMed:8809760] [WorldCat.org] [DOI] (P p)
