NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02999
- pan locus tag?: SAUPAN006408000
- symbol: SAOUHSC_02999
- pan gene symbol?: cap1B
- synonym:
- product: capsular polysaccharide biosynthesis protein Cap5B
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02999
- symbol: SAOUHSC_02999
- product: capsular polysaccharide biosynthesis protein Cap5B
- replicon: chromosome
- strand: -
- coordinates: 2772230..2772922
- length: 693
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921481 NCBI
- RefSeq: YP_501448 NCBI
- BioCyc: G1I0R-2821 BioCyc
- MicrobesOnline: 1291419 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661ATGACGAATACACGAAGAAGTACATCAAGTTTAATTGTCCATGAACAACCAAAGTCACCT
ATTAGCGAGAAATTTCGAGGCATAAGATCAAATATTATGTTTGCAAATCCTGACAGTGCA
GTTCAAAGCATTGTAATCACTTCAGAGGCACCAGGCGCAGGTAAGTCTACAATTGCAGCA
AATTTAGCAGTTGCATATGCGCAAGCAGGTTATAAAACACTAATCGTAGACGGGGATATG
CGTAAACCTACGCAGCATTATATTTTTAATTTGCCAAACAATGAAGGCCTATCAAGTTTA
TTGCTAAATTGGTCAACTTATCAAGACAGTATTATCTCAACTGAAATTCAAGATTTAGAC
GTCTTGACGTCTGGGCCAATCCCACCGAATCCGTCAGAGTTAATTACATCAAGGGCATTT
GCAAATTTGTATGACACATTATTGATGAATTATAACTTTGTAATTATCGATACGCCACCA
GTGAACACAGTTACAGATGCGCAATTATTTTCAAAGTTTACCGGCAATGTTGTCTACGTA
GTTAATTCGGAAAATAATAATAGAGATGAAGTTAAAAAAGGAAAAGAACTTATTGAAGCA
ACAGGTGCTAAATTATTAGGTGTAGTCTTAAATAGAATGCCTAAAGATAAAAGTGCTAGT
TACTATGCATATTATGGGACTGATGAATCATGA60
120
180
240
300
360
420
480
540
600
660
693
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02999
- symbol: SAOUHSC_02999
- description: capsular polysaccharide biosynthesis protein Cap5B
- length: 230
- theoretical pI: 5.30455
- theoretical MW: 25251.3
- GRAVY: -0.247391
⊟Function[edit | edit source]
- TIGRFAM: Transport and binding proteins Carbohydrates, organic alcohols, and acids capsular exopolysaccharide family (TIGR01007; HMM-score: 258.7)and 17 morechain length determinant protein tyrosine kinase EpsG (TIGR03029; HMM-score: 145.5)exopolysaccharide/PEP-CTERM locus tyrosine autokinase (TIGR03018; EC 2.7.10.2; HMM-score: 129.1)Transport and binding proteins Carbohydrates, organic alcohols, and acids exopolysaccharide transport protein family (TIGR01005; HMM-score: 113.1)cell division ATPase MinD (TIGR01969; HMM-score: 55.5)Cellular processes Cell division septum site-determining protein MinD (TIGR01968; HMM-score: 40.2)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides cellulose synthase operon protein YhjQ (TIGR03371; HMM-score: 37.4)helicase/secretion neighborhood CpaE-like protein (TIGR03815; HMM-score: 25.7)Central intermediary metabolism Nitrogen fixation nitrogenase iron protein (TIGR01287; EC 1.18.6.1; HMM-score: 17.4)Mobile and extrachromosomal element functions Plasmid functions plasmid partitioning protein RepA (TIGR03453; HMM-score: 17.4)Hypothetical proteins Conserved transport-energizing ATPase, TRC40/GET3/ArsA family (TIGR00345; EC 3.6.1.-; HMM-score: 16.7)arsenical pump-driving ATPase (TIGR04291; EC 3.6.1.-; HMM-score: 16.3)Central intermediary metabolism Sulfur metabolism adenylyl-sulfate kinase (TIGR00455; EC 2.7.1.25; HMM-score: 16.2)putative cytidylate kinase (TIGR02173; EC 2.7.4.14; HMM-score: 14.1)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll light-independent protochlorophyllide reductase, iron-sulfur ATP-binding protein (TIGR01281; EC 1.3.7.7; HMM-score: 13.8)signal recognition particle protein SRP54 (TIGR01425; HMM-score: 12.8)Protein fate Protein and peptide secretion and trafficking signal recognition particle-docking protein FtsY (TIGR00064; HMM-score: 11.7)HprK-related kinase A (TIGR04352; HMM-score: 10.9)
- TheSEED :
- Capsular polysaccharide synthesis enzyme Cap5B
- Tyrosine-protein kinase EpsD (EC 2.7.10.2)
Cell Wall and Capsule Capsular and extracellular polysacchrides Exopolysaccharide Biosynthesis Tyrosine-protein kinase EpsD (EC 2.7.10.2)and 1 more - PFAM: P-loop_NTPase (CL0023) CbiA; CobQ/CobB/MinD/ParA nucleotide binding domain (PF01656; HMM-score: 51.7)ParA; NUBPL iron-transfer P-loop NTPase (PF10609; HMM-score: 46.5)AAA_31; AAA domain (PF13614; HMM-score: 45.8)and 14 moreMipZ; ATPase MipZ (PF09140; HMM-score: 35.6)CBP_BcsQ; Cellulose biosynthesis protein BcsQ (PF06564; HMM-score: 27.7)ArsA_ATPase; Anion-transporting ATPase (PF02374; HMM-score: 20.1)AAA_26; AAA domain (PF13500; HMM-score: 20)AAA_25; AAA domain (PF13481; HMM-score: 16.9)APS_kinase; Adenylylsulphate kinase (PF01583; HMM-score: 16.4)CLP1_P; mRNA cleavage and polyadenylation factor CLP1 P-loop (PF16575; HMM-score: 15.9)SRP54; SRP54-type protein, GTPase domain (PF00448; HMM-score: 14.8)Fer4_NifH; 4Fe-4S iron sulfur cluster binding proteins, NifH/frxC family (PF00142; HMM-score: 14.6)KTI12; Chromatin associated protein KTI12 (PF08433; HMM-score: 12.3)AAA_29; P-loop containing region of AAA domain (PF13555; HMM-score: 12.2)ArgK; ArgK protein (PF03308; HMM-score: 12)AAA_24; AAA domain (PF13479; HMM-score: 12)AAA_30; AAA domain (PF13604; HMM-score: 12)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0.01
- Cytoplasmic Membrane Score: 9.99
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 0
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.024914
- TAT(Tat/SPI): 0.011712
- LIPO(Sec/SPII): 0.002336
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTNTRRSTSSLIVHEQPKSPISEKFRGIRSNIMFANPDSAVQSIVITSEAPGAGKSTIAANLAVAYAQAGYKTLIVDGDMRKPTQHYIFNLPNNEGLSSLLLNWSTYQDSIISTEIQDLDVLTSGPIPPNPSELITSRAFANLYDTLLMNYNFVIIDTPPVNTVTDAQLFSKFTGNVVYVVNSENNNRDEVKKGKELIEATGAKLLGVVLNRMPKDKSASYYAYYGTDES
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
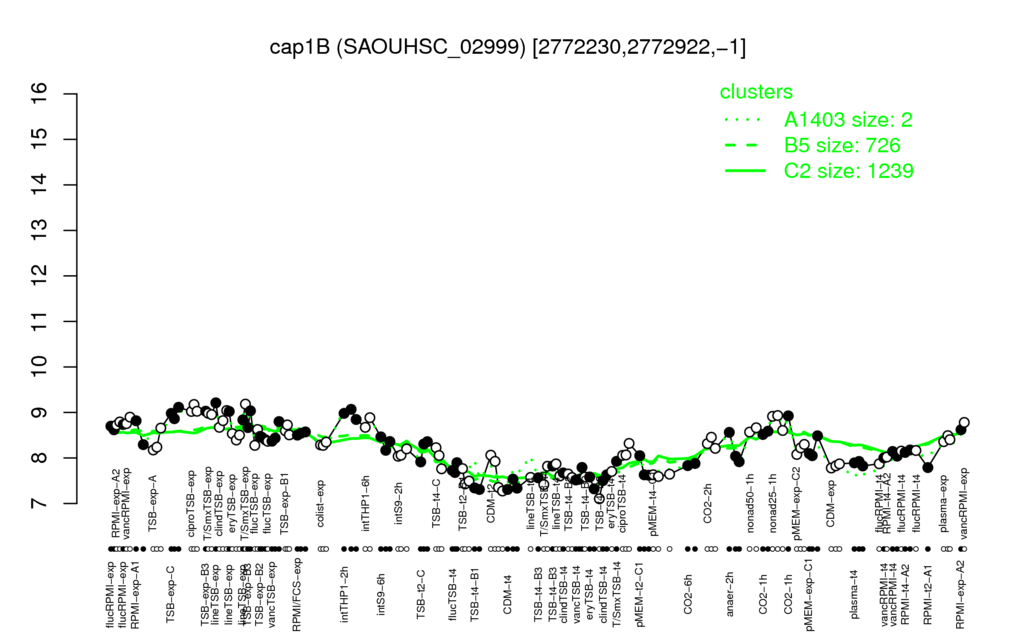 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
