Jump to navigation
Jump to search
m (Text replacement - "<protect> =Summary= * <aureodatabase>organism" to "<protect> <aureodatabase>NCBI date</aureodatabase> =Summary= * <aureodatabase>organism") |
m (Text replacement - "* <aureodatabase>protein Genbank</aureodatabase> " to "") |
||
| Line 101: | Line 101: | ||
* <aureodatabase>protein GI</aureodatabase> | * <aureodatabase>protein GI</aureodatabase> | ||
* <aureodatabase>protein UniProt</aureodatabase> | * <aureodatabase>protein UniProt</aureodatabase> | ||
* <aureodatabase>protein RefSeq</aureodatabase> | * <aureodatabase>protein RefSeq</aureodatabase> | ||
</protect> | </protect> | ||
Revision as of 16:46, 10 March 2016
NCBI date : _
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02595
- pan locus tag?: SAUPAN005801000
- symbol: SAOUHSC_02595
- pan gene symbol?: —
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02595
- symbol: SAOUHSC_02595
- product: hypothetical protein
- replicon: chromosome
- strand: -
- coordinates: 2384973..2385890
- length: 918
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921591 NCBI
- gene Genbank : _
⊟Phenotype[edit | edit source]
- Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901ATGTTTCAAAAATTGAGTATATTTGCCACGAAGAGTTTTTTGGTATGGATGTTAGTAGCA
GCTGTTATTGGATTTATTTTCCCACAACATGTTGCGACATTAGGTAAATGGGTACCTTAT
TTACTTGGTATTGTTATGTTAGGTATGGGATTAACAATTACACCTAATGATTTCAAAATG
GTCTTTAAAGCACCTAGAGCAGTAATTATTGGTGTCTGTCTACAATTCAGTATTATGCCC
ACATTAGCATTTATAATTGCAAAGTCTTTTCATTTACCACCTGATATTGCTGTTGGCGTA
ATATTAGTTGGATGTTGTCCGGGTGGGACATCAAGTAATGTAATGAGTTATTTAGCCAAA
GCTAACGTAGCACTTTCTGTTTCTATTACGACGGTCTCTACGTTGCTAGCGCCATTCGTT
ACACCTGCGTTAATATATCTATTTGCAAATGAATGGTTGGAAGTATCTTTCGTGAGTATG
TTGTGGTCAGTTGTTCAAGTTGTATTAATTCCAATTGCTTTAGGTATTGTTTTGCAAATT
ATTAATCGTAAAATTGCTGAAAAAGCTTCTACAGCTTTGCCAATTATATCAGTTGTTGCT
ATTTCATTAATTTTAGCAATAGTTGTAGGTGGCAGTAAGCACCAAATCTTAACTACAGGA
TTATTAATATTTTTAGTAGTTATTTTACATAACGTATTAGGGTATACGATTGGATATTGG
TTAGCTCGTCTTTTAAAATTAGATCGACAAGATCAAAAAGCAGTCAGTATTGAAGTTGGA
ATGCAGAACTCTGGTTTAGCTGTGTCATTAGCAGCATTGCATTTTAATCCAATTGCAGCA
GTACCAGGCGCAGTGTTTAGTTTCATTCATAATATAACAGGGCCTATTTTAGCAAAGTAT
TGGTCAAAAAAGTTATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
918
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02595
- symbol: SAOUHSC_02595
- description: hypothetical protein
- length: 305
- theoretical pI: 10.2753
- theoretical MW: 32851.6
- GRAVY: 1.06951
⊟Function[edit | edit source]
- TIGRFAM: Transport and binding proteins Carbohydrates, organic alcohols, and acids bile acid transporter (TIGR00841; HMM-score: 289)and 1 moremembrane protein AbrB duplication (TIGR03082; HMM-score: 11.3)
- TheSEED :
- COG0385 sodium-dependent transporter
- PFAM: CPA_AT (CL0064) SBF; Sodium Bile acid symporter family (PF01758; HMM-score: 155.2)and 3 moreSBF_like; SBF-like CPA transporter family (DUF4137) (PF13593; HMM-score: 77.2)Mem_trans; Membrane transport protein (PF03547; HMM-score: 10.4)no clan defined DUF3464; Protein of unknown function (DUF3464) (PF11947; HMM-score: 6.3)
⊟Structure, modifications & interactions[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
- interaction partners:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 8
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: 0
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.025516
- TAT(Tat/SPI): 0.001144
- LIPO(Sec/SPII): 0.015437
- predicted transmembrane helices (TMHMM): 9
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MFQKLSIFATKSFLVWMLVAAVIGFIFPQHVATLGKWVPYLLGIVMLGMGLTITPNDFKMVFKAPRAVIIGVCLQFSIMPTLAFIIAKSFHLPPDIAVGVILVGCCPGGTSSNVMSYLAKANVALSVSITTVSTLLAPFVTPALIYLFANEWLEVSFVSMLWSVVQVVLIPIALGIVLQIINRKIAEKASTALPIISVVAISLILAIVVGGSKHQILTTGLLIFLVVILHNVLGYTIGYWLARLLKLDRQDQKAVSIEVGMQNSGLAVSLAALHFNPIAAVPGAVFSFIHNITGPILAKYWSKKL
⊟Peptides[edit | edit source]
- experimentally validated:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [1] : SAOUHSC_02593 < SAOUHSC_02594 < SAOUHSC_02595
⊟Regulation[edit | edit source]
- sigma factors : _
- regulator: CodY* (repression) regulon
CodY* (TF) important in Amino acid metabolism; RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
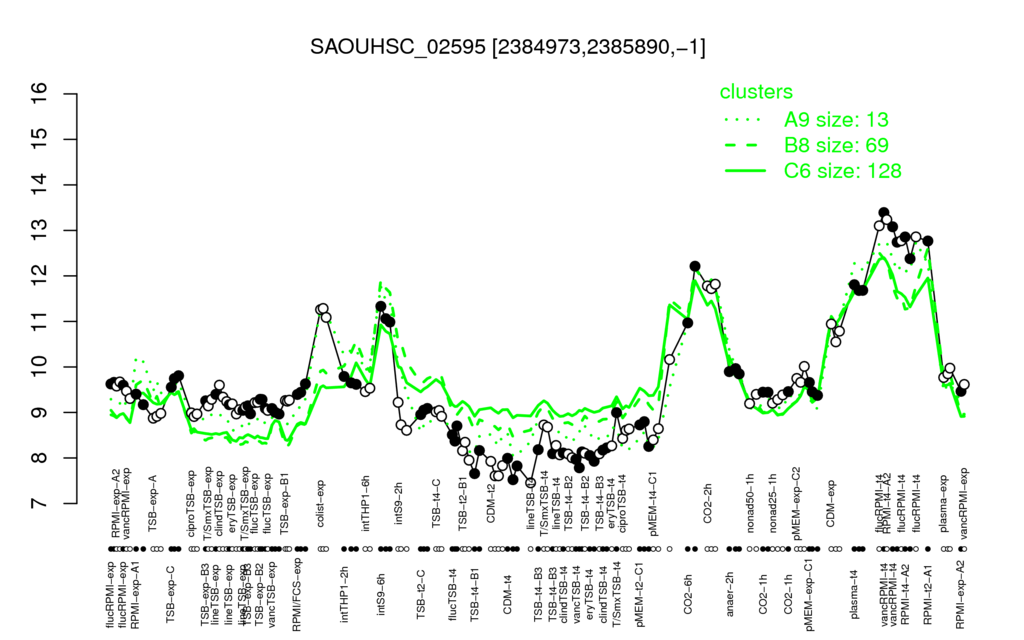 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ 1.0 1.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
