Jump to navigation
Jump to search
m (Text replacement - "* <aureodatabase>protein Genbank</aureodatabase> " to "") |
m (Text replacement - "gene Genbank" to "gene RefSeq") |
||
| Line 1: | Line 1: | ||
__TOC__ | |||
<protect> | <protect> | ||
<aureodatabase> | <aureodatabase>annotation</aureodatabase> | ||
=Summary= | =Summary= | ||
* <aureodatabase>organism</aureodatabase> | *<aureodatabase>organism</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>pan locus</aureodatabase> | *<aureodatabase>pan locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>pan gene symbol</aureodatabase> | *<aureodatabase>pan gene symbol</aureodatabase> | ||
* <aureodatabase>gene synonyms</aureodatabase> | *<aureodatabase>gene synonyms</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
</protect> | </protect> | ||
| Line 24: | Line 25: | ||
==General== | ==General== | ||
* <aureodatabase>gene type</aureodatabase> | *<aureodatabase>gene type</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
* <aureodatabase>gene replicon</aureodatabase> | *<aureodatabase>gene replicon</aureodatabase> | ||
* <aureodatabase>strand</aureodatabase> | *<aureodatabase>strand</aureodatabase> | ||
* <aureodatabase>gene coordinates</aureodatabase> | *<aureodatabase>gene coordinates</aureodatabase> | ||
* <aureodatabase>gene length</aureodatabase> | *<aureodatabase>gene length</aureodatabase> | ||
* <aureodatabase>essential</aureodatabase> | *<aureodatabase>essential</aureodatabase> | ||
*<aureodatabase>gene comment</aureodatabase> | |||
</protect> | </protect> | ||
| Line 38: | Line 40: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>gene GI</aureodatabase> | *<aureodatabase>gene GI</aureodatabase> | ||
* <aureodatabase>gene | *<aureodatabase>gene RefSeq</aureodatabase> | ||
*<aureodatabase>gene BioCyc</aureodatabase> | |||
*<aureodatabase>gene MicrobesOnline</aureodatabase> | |||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Phenotype== | ==Phenotype== | ||
</protect> | </protect> | ||
Share your knowledge and add information here. [<span class="plainlinks">[//aureowiki.med.uni-greifswald.de/index.php?title={{PAGENAMEE}}&veaction=edit§ion=6 edit]</span>] | |||
<protect> | <protect> | ||
==DNA sequence== | ==DNA sequence== | ||
* <aureodatabase>gene sequence</aureodatabase> | *<aureodatabase>gene sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
<aureodatabase>RNA regulated operons</aureodatabase> | |||
</protect> | |||
<protect> | |||
=Protein= | =Protein= | ||
<aureodatabase>protein 3D view</aureodatabase> | <aureodatabase>protein 3D view</aureodatabase> | ||
==General== | ==General== | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>protein symbol</aureodatabase> | *<aureodatabase>protein symbol</aureodatabase> | ||
* <aureodatabase>protein description</aureodatabase> | *<aureodatabase>protein description</aureodatabase> | ||
* <aureodatabase>protein length</aureodatabase> | *<aureodatabase>protein length</aureodatabase> | ||
* <aureodatabase>theoretical pI</aureodatabase> | *<aureodatabase>theoretical pI</aureodatabase> | ||
* <aureodatabase>theoretical MW</aureodatabase> | *<aureodatabase>theoretical MW</aureodatabase> | ||
* <aureodatabase>GRAVY</aureodatabase> | *<aureodatabase>GRAVY</aureodatabase> | ||
</protect> | </protect> | ||
| Line 71: | Line 78: | ||
==Function== | ==Function== | ||
* <aureodatabase>protein reaction</aureodatabase> | *<aureodatabase>protein reaction</aureodatabase> | ||
* <aureodatabase>protein TIGRFAM</aureodatabase> | *<aureodatabase>protein TIGRFAM</aureodatabase> | ||
* <aureodatabase>protein TheSeed</aureodatabase> | *<aureodatabase>protein TheSeed</aureodatabase> | ||
* <aureodatabase>protein PFAM</aureodatabase> | *<aureodatabase>protein PFAM</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Structure, modifications & | ==Structure, modifications & cofactors== | ||
* <aureodatabase>protein domains</aureodatabase> | *<aureodatabase>protein domains</aureodatabase> | ||
* <aureodatabase>protein modifications</aureodatabase> | *<aureodatabase>protein modifications</aureodatabase> | ||
* <aureodatabase>protein cofactors</aureodatabase> | *<aureodatabase>protein cofactors</aureodatabase> | ||
* <aureodatabase>protein effectors</aureodatabase> | *<aureodatabase>protein effectors</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein regulated operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 90: | Line 97: | ||
==Localization== | ==Localization== | ||
* <aureodatabase>protein Psortb</aureodatabase> | *<aureodatabase>protein Psortb</aureodatabase> | ||
* <aureodatabase>protein LocateP</aureodatabase> | *<aureodatabase>protein LocateP</aureodatabase> | ||
* <aureodatabase>protein SignalP</aureodatabase> | *<aureodatabase>protein SignalP</aureodatabase> | ||
* <aureodatabase>protein TMHMM</aureodatabase> | *<aureodatabase>protein TMHMM</aureodatabase> | ||
</protect> | </protect> | ||
| Line 99: | Line 106: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>protein GI</aureodatabase> | *<aureodatabase>protein GI</aureodatabase> | ||
* <aureodatabase>protein UniProt</aureodatabase> | *<aureodatabase>protein RefSeq</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein UniProt</aureodatabase> | ||
*<aureodatabase>protein STRING</aureodatabase> | |||
</protect> | </protect> | ||
| Line 107: | Line 115: | ||
==Protein sequence== | ==Protein sequence== | ||
* <aureodatabase>protein sequence</aureodatabase> | *<aureodatabase>protein sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Experimental data== | ||
* <aureodatabase>protein validated peptides</aureodatabase> | *<aureodatabase>protein validated peptides</aureodatabase> | ||
*<aureodatabase>protein validated localization</aureodatabase> | |||
*<aureodatabase>protein validated quantitative data</aureodatabase> | |||
*<aureodatabase>protein partners</aureodatabase> | |||
</protect> | </protect> | ||
| Line 124: | Line 135: | ||
==Operon== | ==Operon== | ||
* <aureodatabase>operons</aureodatabase> | *<aureodatabase>operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 130: | Line 141: | ||
==Regulation== | ==Regulation== | ||
*<aureodatabase>regulators</aureodatabase> | |||
* <aureodatabase>regulators</aureodatabase> | |||
</protect> | </protect> | ||
| Line 137: | Line 147: | ||
==Transcription pattern== | ==Transcription pattern== | ||
* <aureodatabase>expression browser</aureodatabase> | *<aureodatabase>expression browser</aureodatabase> | ||
</protect> | </protect> | ||
| Line 143: | Line 153: | ||
==Protein synthesis (provided by Aureolib)== | ==Protein synthesis (provided by Aureolib)== | ||
* <aureodatabase>protein synthesis Aureolib</aureodatabase> | *<aureodatabase>protein synthesis Aureolib</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Protein stability== | ||
* <aureodatabase>protein half-life</aureodatabase> | *<aureodatabase>protein half-life</aureodatabase> | ||
</protect> | </protect> | ||
Latest revision as of 12:27, 11 March 2016
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01383
- pan locus tag?: SAUPAN003790000
- symbol: SAOUHSC_01383
- pan gene symbol?: —
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_01383
- symbol: SAOUHSC_01383
- product: hypothetical protein
- replicon: chromosome
- strand: +
- coordinates: 1325827..1327641
- length: 1815
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3920792 NCBI
- RefSeq: YP_499910 NCBI
- BioCyc: G1I0R-1293 BioCyc
- MicrobesOnline: 1289824 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801ATGTCTCAAGGTTTACCTTTAAGAGAAGATGTTCCTGTTTCAGAAACATGGGATTTAGTA
GACTTATTTAAAGATGATCAACAATATTATGAAAGTATTGACGCTCTAGTACAACAAGCA
AATCAATTTCATCATACATATGCAACAACATTAAATTCAATCGAACAAATTAATACTGCT
TTAGCTGAATTAGAAAATATTTTAATTGCCTTAGATCGCTTAAGTAATTATGCAGAACTA
CGTTTAAGTGTAGATACTAGTAATATCGAGGCACAAGTATTGAGCGCTAAATTATCTACT
ACATACGGTAAAATTGTTAGCCAATTATCATTTGTAGAGTCAGAAATACTTGAATTACCA
GAAGAAATACTTCAACAATTAGAAGAATCATGTCCATATCAACACTATATTAAACAGTTA
ATAAAACAAAAGCCATTCCAATTATCTGCGTCGGTAGAACAAGTATTAGCAACTTTATCA
CCTACGCTAAACAGTCCTTACGATTTATACGGCACGACAAAAATGCTAGATATTACATTC
GATTCATTTGAACATGATGGTACAACGTACCCTGTCGACTATGCTACGTTTGAAAATGAT
TATGAAGATAATAAAGATCCTGAGTTTAGACGTAAAAGTTTCAAATCGTTTAGCGATGGG
ATTCGAAAATATCAGCATACTACCGCGGCTACATATAATATGCAAGTACAACAAGAAAAA
ATTGAAGCTGATTTACGTGGATTTGAATCAGTCATCGATTATTTATTACATAGTCAAGAA
GTAACGCGTGATATGTTTGACCGTCAAATCGATATGATTATGCGTGACTTGGCACCAGTT
ATGCAGAAATATGCTAAACTTTTACAACGTATTCACGGATTAGATAACATGCGTTTTGAA
GACTTGAAGATTTCTGTAGACCCTGATTATGAACCAGAGATTTCAATTGAAGACTCAAAA
AATTATATTTTCGGTGCGTTAAGTGTTTTAGGTGATGACTATACAAACATGTTACGTGAA
GCATACGATCAGCGATGGATTGATTTTGCACAAAATAAAGGTAAAGATACAGGCGCATTT
TGTGCAAGTCCATACTTTACACATTCATATGTGTTTATTTCTTGGACTGGTAAAATGGCT
GAAGCATTTGTCTTAGCACATGAATTAGGTCATGCAGGTCATTTTACATTAGCTCAAAAA
CATCAACCATATCTTGAATCAGAAGCATCAATGTACTTTGTTGAAGCCCCTTCTACAATG
AATGAAATGTTGATGGCCAATTATTTATTTAACACAAGTGATAATCCAAGATTTAAGCGT
TGGGTTATTGGCTCAATTTTATCTAGAACATATTATCATAATATGGTTACCCATTTATTA
GAAGCTGCTTATCAACGTGAAGTGTATCACAAAGTAGATCAAGGTGAATCTTTAAATGCG
CCGACATTAAATGAAATAATGCTAAATGTTTATAAACAATTTTTTGGAGATGCAGTAGAC
ATGACTGAGGGTGCTGAATTAACATGGATGCGTCAACCTCATTACTATATGGGATTATAT
TCGTATACGTATTCTGCTGGCTTAACAATCGGAACTGTCGTTTCTCAAAAGATTAAAAAT
GAAGGCCAACCAGCTGTTGATGCTTGGTTAGAAACATTGAAAAAAGGTGGTAGTGTATCA
CCTGTCGAACTTGCAAACATTGCAGGTGTAGACATTACTACAGAACAGCCACTTAAATCT
ACAATTCAATATATTTCTGATTTAGTCGATGAAGTTGAAAAATTAACAGATGAAATTGAG
CAAGCAAATAACTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1815
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_01383
- symbol: SAOUHSC_01383
- description: hypothetical protein
- length: 604
- theoretical pI: 4.38757
- theoretical MW: 69284.2
- GRAVY: -0.402152
⊟Function[edit | edit source]
- TIGRFAM: Protein fate Degradation of proteins, peptides, and glycopeptides oligoendopeptidase F (TIGR00181; EC 3.4.24.-; HMM-score: 720.9)and 2 moreoligoendopeptidase, pepF/M3 family (TIGR02290; EC 3.4.24.-; HMM-score: 282.3)oligoendopeptidase, M3 family (TIGR02289; EC 3.4.24.-; HMM-score: 66.6)
- TheSEED :
- Oligoendopeptidase F-like protein
- PFAM: Peptidase_MA (CL0126) Peptidase_M3; Peptidase family M3 (PF01432; HMM-score: 123.3)and 4 moreno clan defined Peptidase_M3_N; Oligopeptidase F (PF08439; HMM-score: 54.1)BLOC1_2; Biogenesis of lysosome-related organelles complex-1 subunit 2 (PF10046; HMM-score: 8.9)LOH1CR12; Tumour suppressor protein (PF10158; HMM-score: 8.3)EsxAB (CL0352) WXG100; Proteins of 100 residues with WXG (PF06013; HMM-score: 6.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: Zn2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.001997
- TAT(Tat/SPI): 0.000547
- LIPO(Sec/SPII): 0.000272
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSQGLPLREDVPVSETWDLVDLFKDDQQYYESIDALVQQANQFHHTYATTLNSIEQINTALAELENILIALDRLSNYAELRLSVDTSNIEAQVLSAKLSTTYGKIVSQLSFVESEILELPEEILQQLEESCPYQHYIKQLIKQKPFQLSASVEQVLATLSPTLNSPYDLYGTTKMLDITFDSFEHDGTTYPVDYATFENDYEDNKDPEFRRKSFKSFSDGIRKYQHTTAATYNMQVQQEKIEADLRGFESVIDYLLHSQEVTRDMFDRQIDMIMRDLAPVMQKYAKLLQRIHGLDNMRFEDLKISVDPDYEPEISIEDSKNYIFGALSVLGDDYTNMLREAYDQRWIDFAQNKGKDTGAFCASPYFTHSYVFISWTGKMAEAFVLAHELGHAGHFTLAQKHQPYLESEASMYFVEAPSTMNEMLMANYLFNTSDNPRFKRWVIGSILSRTYYHNMVTHLLEAAYQREVYHKVDQGESLNAPTLNEIMLNVYKQFFGDAVDMTEGAELTWMRQPHYYMGLYSYTYSAGLTIGTVVSQKIKNEGQPAVDAWLETLKKGGSVSPVELANIAGVDITTEQPLKSTIQYISDLVDEVEKLTDEIEQANN
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
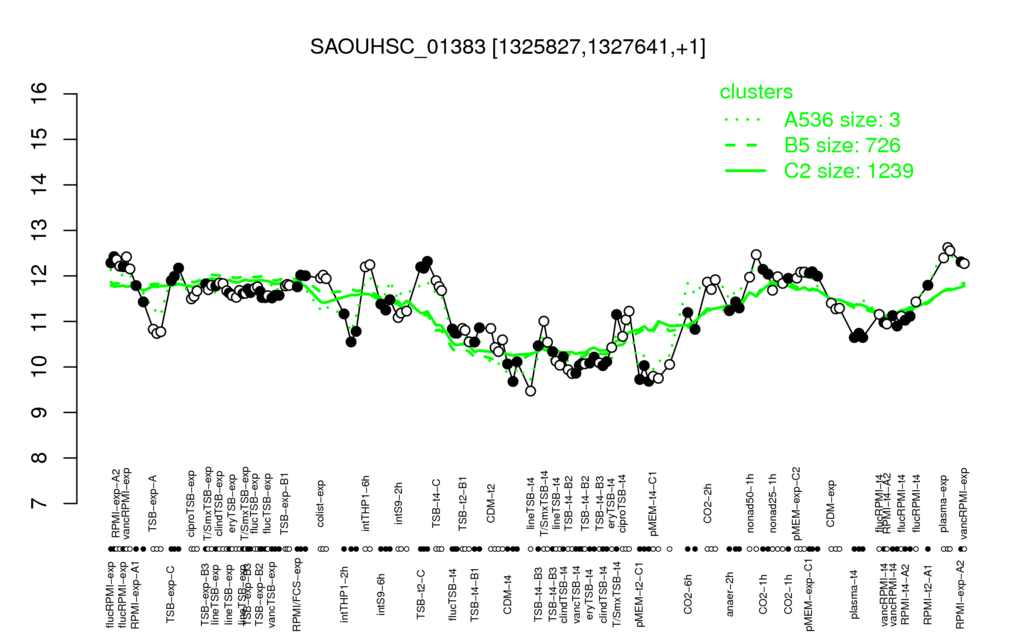 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
