⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00365
- pan locus tag?: SAUPAN001982000
- symbol: SAOUHSC_00365
- pan gene symbol?: ahpC
- synonym:
- product: alkyl hydroperoxide reductase subunit C
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00365
- symbol: SAOUHSC_00365
- product: alkyl hydroperoxide reductase subunit C
- replicon: chromosome
- strand: -
- coordinates: 371268..371837
- length: 570
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
⊟Phenotype[edit | edit source]
- Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541ATGTCATTAATTAACAAAGAAATCTTACCATTTACAGCGCAAGCTTTCGATCCAAAAAAA
GATCAATTTAAAGAAGTTACACAAGAAGATTTAAAAGGTTCTTGGAGCGTAGTATGCTTC
TATCCTGCTGACTTCTCATTCGTTTGTCCAACTGAATTAGAAGACTTACAAAACCAATAT
GAAGAATTACAAAAATTAGGCGTAAATGTATTCTCAGTATCAACTGATACTCACTTCGTA
CACAAAGCATGGCATGACCATTCAGATGCAATTAGCAAAATCACTTACACTATGATTGGT
GACCCATCACAAACAATCACTCGTAATTTTGATGTATTAGATGAAGCTACTGGTTTAGCT
CAACGTGGTACATTCATTATCGACCCAGACGGTGTTGTACAAGCATCTGAAATTAACGCT
GACGGAATTGGCCGTGACGCTAGTACATTAGCTCACAAAATCAAAGCAGCTCAATATGTT
CGTAAAAACCCTGGCGAAGTATGCCCAGCTAAATGGGAAGAAGGCGCTAAAACATTGCAA
CCTGGTTTAGATTTAGTAGGTAAAATCTAA60
120
180
240
300
360
420
480
540
570
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00365
- symbol: SAOUHSC_00365
- description: alkyl hydroperoxide reductase subunit C
- length: 189
- theoretical pI: 4.6567
- theoretical MW: 20976.5
- GRAVY: -0.331217
⊟Function[edit | edit source]
- reaction: EC 1.11.1.15? ExPASyPeroxiredoxin 2 R'-SH + ROOH = R'-S-S-R' + H2O + ROH
- TIGRFAM: Cellular processes Detoxification peroxiredoxin (TIGR03137; EC 1.11.1.15; HMM-score: 321.6)Cellular processes Adaptations to atypical conditions peroxiredoxin (TIGR03137; EC 1.11.1.15; HMM-score: 321.6)
- TheSEED :
- NADH-dependent alkyl hydroperoxide reductase component AhpC (EC 1.11.1.26)
- PFAM: Thioredoxin (CL0172) AhpC-TSA; AhpC/TSA family (PF00578; HMM-score: 119.2)and 4 moreRedoxin; Redoxin (PF08534; HMM-score: 52.2)no clan defined 1-cysPrx_C; C-terminal domain of 1-Cys peroxiredoxin (PF10417; HMM-score: 25)Thioredoxin (CL0172) SCO1-SenC; SCO1/SenC (PF02630; HMM-score: 21.2)Glyco_hydro_tim (CL0058) NAGidase; beta-N-acetylglucosaminidase (PF07555; HMM-score: 12.6)
⊟Structure, modifications & interactions[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
- interaction partners:
SAOUHSC_01170 (carB) carbamoyl phosphate synthase large subunit [1] (data from MRSA252) SAOUHSC_01683 (dnaK) molecular chaperone DnaK [1] (data from MRSA252) SAOUHSC_00799 (eno) phosphopyruvate hydratase [1] (data from MRSA252) SAOUHSC_02117 (gatA) aspartyl/glutamyl-tRNA amidotransferase subunit A [1] (data from MRSA252) SAOUHSC_02254 (groEL) chaperonin GroEL [1] (data from MRSA252) SAOUHSC_00519 (rplA) 50S ribosomal protein L1 [1] (data from MRSA252) SAOUHSC_02509 (rplB) 50S ribosomal protein L2 [1] (data from MRSA252) SAOUHSC_02512 (rplC) 50S ribosomal protein L3 [1] (data from MRSA252) SAOUHSC_02511 (rplD) 50S ribosomal protein L4 [1] (data from MRSA252) SAOUHSC_02496 (rplF) 50S ribosomal protein L6 [1] (data from MRSA252) SAOUHSC_00520 (rplJ) 50S ribosomal protein L10 [1] (data from MRSA252) SAOUHSC_00521 (rplL) 50S ribosomal protein L7/L12 [1] (data from MRSA252) SAOUHSC_02492 (rplO) 50S ribosomal protein L15 [1] (data from MRSA252) SAOUHSC_02505 (rplP) 50S ribosomal protein L16 [1] (data from MRSA252) SAOUHSC_02495 (rplR) 50S ribosomal protein L18 [1] (data from MRSA252) SAOUHSC_01211 (rplS) 50S ribosomal protein L19 [1] (data from MRSA252) SAOUHSC_01784 (rplT) 50S ribosomal protein L20 [1] (data from MRSA252) SAOUHSC_01757 (rplU) 50S ribosomal protein L21 [1] (data from MRSA252) SAOUHSC_02507 (rplV) 50S ribosomal protein L22 [1] (data from MRSA252) SAOUHSC_02510 (rplW) 50S ribosomal protein L23 [1] (data from MRSA252) SAOUHSC_01232 (rpsB) 30S ribosomal protein S2 [1] (data from MRSA252) SAOUHSC_01829 (rpsD) 30S ribosomal protein S4 [1] (data from MRSA252) SAOUHSC_02494 (rpsE) 30S ribosomal protein S5 [1] (data from MRSA252) SAOUHSC_02477 (rpsI) 30S ribosomal protein S9 [1] (data from MRSA252) SAOUHSC_02503 (rpsQ) 30S ribosomal protein S17 [1] (data from MRSA252) SAOUHSC_02508 (rpsS) 30S ribosomal protein S19 [1] (data from MRSA252) SAOUHSC_01418 (sucA) 2-oxoglutarate dehydrogenase E1 component [1] (data from MRSA252) SAOUHSC_00187 formate acetyltransferase [1] (data from MRSA252) SAOUHSC_00529 elongation factor G [1] (data from MRSA252) SAOUHSC_00530 elongation factor Tu [1] (data from MRSA252) SAOUHSC_00679 hypothetical protein [1] (data from MRSA252) SAOUHSC_00878 hypothetical protein [1] (data from MRSA252) SAOUHSC_00947 enoyl-(acyl carrier protein) reductase [1] (data from MRSA252) SAOUHSC_01040 pyruvate dehydrogenase complex, E1 component subunit alpha [1] (data from MRSA252) SAOUHSC_01150 cell division protein FtsZ [1] (data from MRSA252) SAOUHSC_01490 DNA-binding protein HU [1] (data from MRSA252) SAOUHSC_01801 isocitrate dehydrogenase [1] (data from MRSA252) SAOUHSC_01806 pyruvate kinase [1] (data from MRSA252) SAOUHSC_01819 hypothetical protein [1] (data from MRSA252) SAOUHSC_02377 pyrimidine-nucleoside phosphorylase [1] (data from MRSA252) SAOUHSC_02441 alkaline shock protein 23 [1] (data from MRSA252) SAOUHSC_02485 DNA-directed RNA polymerase subunit alpha [1] (data from MRSA252) SAOUHSC_02486 30S ribosomal protein S11 [1] (data from MRSA252) SAOUHSC_02849 pyruvate oxidase [1] (data from MRSA252) SAOUHSC_02927 malate:quinone oxidoreductase [1] (data from MRSA252) SAOUHSC_02969 arginine deiminase [1] (data from MRSA252)
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 0
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.012719
- TAT(Tat/SPI): 0.000229
- LIPO(Sec/SPII): 0.000739
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSLINKEILPFTAQAFDPKKDQFKEVTQEDLKGSWSVVCFYPADFSFVCPTELEDLQNQYEELQKLGVNVFSVSTDTHFVHKAWHDHSDAISKITYTMIGDPSQTITRNFDVLDEATGLAQRGTFIIDPDGVVQASEINADGIGRDASTLAHKIKAAQYVRKNPGEVCPAKWEEGAKTLQPGLDLVGKI
⊟Peptides[edit | edit source]
- experimentally validated: PeptideAtlas [2] [3]
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- sigma factors : _
- regulator: PerR* (repression) regulon
PerR* (TF) important in Oxidative stress response; RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
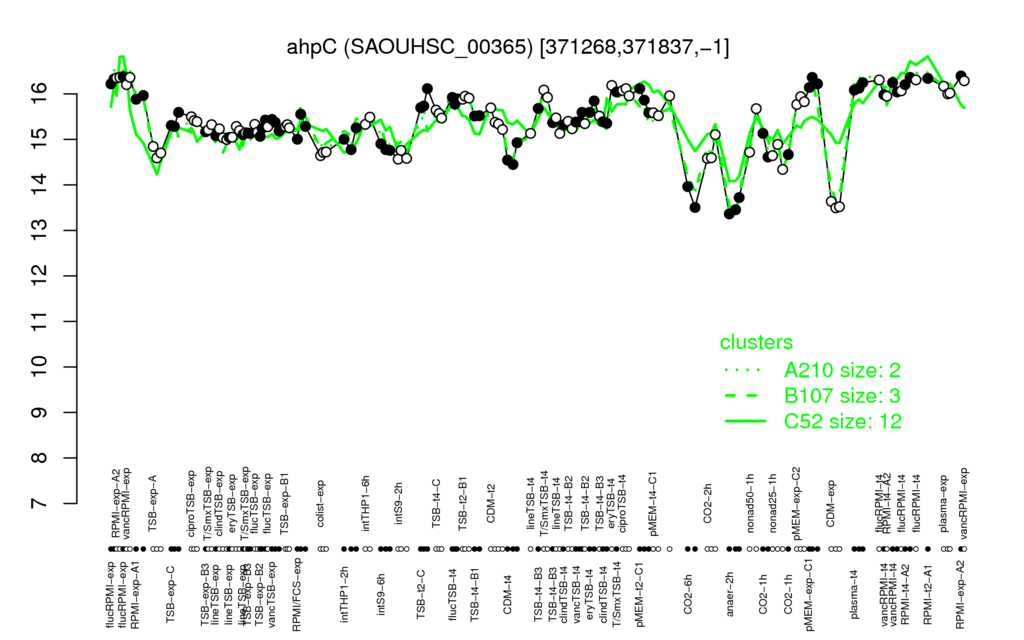 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 1.22 1.23 1.24 1.25 1.26 1.27 1.28 1.29 1.30 1.31 1.32 1.33 1.34 1.35 1.36 1.37 1.38 1.39 1.40 1.41 1.42 1.43 1.44 1.45 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 4.0 4.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
⊟Relevant publications[edit | edit source]
Julie A Morrissey, Alan Cockayne, Kirsty Brummell, Paul Williams
The staphylococcal ferritins are differentially regulated in response to iron and manganese and via PerR and Fur.
Infect Immun: 2004, 72(2);972-9
[PubMed:14742543] [WorldCat.org] [DOI] (P p)Kate Cosgrove, Graham Coutts, Ing-Marie Jonsson, Andrej Tarkowski, John F Kokai-Kun, James J Mond, Simon J Foster
Catalase (KatA) and alkyl hydroperoxide reductase (AhpC) have compensatory roles in peroxide stress resistance and are required for survival, persistence, and nasal colonization in Staphylococcus aureus.
J Bacteriol: 2007, 189(3);1025-35
[PubMed:17114262] [WorldCat.org] [DOI] (P p)L Armstrong-Buisseret, M B Cole, G S Stewart
A homologue to the Escherichia coli alkyl hydroperoxide reductase AhpC is induced by osmotic upshock in Staphylococcus aureus.
Microbiology (Reading): 1995, 141 ( Pt 7);1655-61
[PubMed:7551034] [WorldCat.org] [DOI] (P p)
