Jump to navigation
Jump to search
m (Text replacement - "gene Genbank" to "gene RefSeq") |
m (Text replacement - "* <aureodatabase>protein Genbank</aureodatabase> " to "") |
||
| Line 1: | Line 1: | ||
__TOC__ | |||
<protect> | <protect> | ||
<aureodatabase> | <aureodatabase>annotation</aureodatabase> | ||
=Summary= | =Summary= | ||
* <aureodatabase>organism</aureodatabase> | *<aureodatabase>organism</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>pan locus</aureodatabase> | *<aureodatabase>pan locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>pan gene symbol</aureodatabase> | *<aureodatabase>pan gene symbol</aureodatabase> | ||
* <aureodatabase>gene synonyms</aureodatabase> | *<aureodatabase>gene synonyms</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
</protect> | </protect> | ||
| Line 24: | Line 25: | ||
==General== | ==General== | ||
* <aureodatabase>gene type</aureodatabase> | *<aureodatabase>gene type</aureodatabase> | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>gene symbol</aureodatabase> | *<aureodatabase>gene symbol</aureodatabase> | ||
* <aureodatabase>product</aureodatabase> | *<aureodatabase>product</aureodatabase> | ||
* <aureodatabase>gene replicon</aureodatabase> | *<aureodatabase>gene replicon</aureodatabase> | ||
* <aureodatabase>strand</aureodatabase> | *<aureodatabase>strand</aureodatabase> | ||
* <aureodatabase>gene coordinates</aureodatabase> | *<aureodatabase>gene coordinates</aureodatabase> | ||
* <aureodatabase>gene length</aureodatabase> | *<aureodatabase>gene length</aureodatabase> | ||
* <aureodatabase>essential</aureodatabase> | *<aureodatabase>essential</aureodatabase> | ||
*<aureodatabase>gene comment</aureodatabase> | |||
</protect> | </protect> | ||
| Line 38: | Line 40: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>gene GI</aureodatabase> | *<aureodatabase>gene GI</aureodatabase> | ||
* <aureodatabase>gene RefSeq</aureodatabase> | *<aureodatabase>gene RefSeq</aureodatabase> | ||
*<aureodatabase>gene BioCyc</aureodatabase> | |||
*<aureodatabase>gene MicrobesOnline</aureodatabase> | |||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Phenotype== | ==Phenotype== | ||
</protect> | </protect> | ||
Share your knowledge and add information here. [<span class="plainlinks">[//aureowiki.med.uni-greifswald.de/index.php?title={{PAGENAMEE}}&veaction=edit§ion=6 edit]</span>] | |||
<protect> | <protect> | ||
==DNA sequence== | ==DNA sequence== | ||
* <aureodatabase>gene sequence</aureodatabase> | *<aureodatabase>gene sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
<aureodatabase>RNA regulated operons</aureodatabase> | |||
</protect> | |||
<protect> | |||
=Protein= | =Protein= | ||
<aureodatabase>protein 3D view</aureodatabase> | <aureodatabase>protein 3D view</aureodatabase> | ||
==General== | ==General== | ||
* <aureodatabase>locus</aureodatabase> | *<aureodatabase>locus</aureodatabase> | ||
* <aureodatabase>protein symbol</aureodatabase> | *<aureodatabase>protein symbol</aureodatabase> | ||
* <aureodatabase>protein description</aureodatabase> | *<aureodatabase>protein description</aureodatabase> | ||
* <aureodatabase>protein length</aureodatabase> | *<aureodatabase>protein length</aureodatabase> | ||
* <aureodatabase>theoretical pI</aureodatabase> | *<aureodatabase>theoretical pI</aureodatabase> | ||
* <aureodatabase>theoretical MW</aureodatabase> | *<aureodatabase>theoretical MW</aureodatabase> | ||
* <aureodatabase>GRAVY</aureodatabase> | *<aureodatabase>GRAVY</aureodatabase> | ||
</protect> | </protect> | ||
| Line 71: | Line 78: | ||
==Function== | ==Function== | ||
* <aureodatabase>protein reaction</aureodatabase> | *<aureodatabase>protein reaction</aureodatabase> | ||
* <aureodatabase>protein TIGRFAM</aureodatabase> | *<aureodatabase>protein TIGRFAM</aureodatabase> | ||
* <aureodatabase>protein TheSeed</aureodatabase> | *<aureodatabase>protein TheSeed</aureodatabase> | ||
* <aureodatabase>protein PFAM</aureodatabase> | *<aureodatabase>protein PFAM</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
==Structure, modifications & | ==Structure, modifications & cofactors== | ||
* <aureodatabase>protein domains</aureodatabase> | *<aureodatabase>protein domains</aureodatabase> | ||
* <aureodatabase>protein modifications</aureodatabase> | *<aureodatabase>protein modifications</aureodatabase> | ||
* <aureodatabase>protein cofactors</aureodatabase> | *<aureodatabase>protein cofactors</aureodatabase> | ||
* <aureodatabase>protein effectors</aureodatabase> | *<aureodatabase>protein effectors</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein regulated operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 90: | Line 97: | ||
==Localization== | ==Localization== | ||
* <aureodatabase>protein Psortb</aureodatabase> | *<aureodatabase>protein Psortb</aureodatabase> | ||
* <aureodatabase>protein LocateP</aureodatabase> | *<aureodatabase>protein LocateP</aureodatabase> | ||
* <aureodatabase>protein SignalP</aureodatabase> | *<aureodatabase>protein SignalP</aureodatabase> | ||
* <aureodatabase>protein TMHMM</aureodatabase> | *<aureodatabase>protein TMHMM</aureodatabase> | ||
</protect> | </protect> | ||
| Line 99: | Line 106: | ||
==Accession numbers== | ==Accession numbers== | ||
* <aureodatabase>protein GI</aureodatabase> | *<aureodatabase>protein GI</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein RefSeq</aureodatabase> | ||
* <aureodatabase>protein | *<aureodatabase>protein UniProt</aureodatabase> | ||
</protect> | </protect> | ||
| Line 108: | Line 114: | ||
==Protein sequence== | ==Protein sequence== | ||
* <aureodatabase>protein sequence</aureodatabase> | *<aureodatabase>protein sequence</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Experimental data== | ||
* <aureodatabase>protein validated peptides</aureodatabase> | *<aureodatabase>protein validated peptides</aureodatabase> | ||
*<aureodatabase>protein validated localization</aureodatabase> | |||
*<aureodatabase>protein validated quantitative data</aureodatabase> | |||
*<aureodatabase>protein partners</aureodatabase> | |||
</protect> | </protect> | ||
| Line 125: | Line 134: | ||
==Operon== | ==Operon== | ||
* <aureodatabase>operons</aureodatabase> | *<aureodatabase>operons</aureodatabase> | ||
</protect> | </protect> | ||
| Line 131: | Line 140: | ||
==Regulation== | ==Regulation== | ||
*<aureodatabase>regulators</aureodatabase> | |||
* <aureodatabase>regulators</aureodatabase> | |||
</protect> | </protect> | ||
| Line 138: | Line 146: | ||
==Transcription pattern== | ==Transcription pattern== | ||
* <aureodatabase>expression browser</aureodatabase> | *<aureodatabase>expression browser</aureodatabase> | ||
</protect> | </protect> | ||
| Line 144: | Line 152: | ||
==Protein synthesis (provided by Aureolib)== | ==Protein synthesis (provided by Aureolib)== | ||
* <aureodatabase>protein synthesis Aureolib</aureodatabase> | *<aureodatabase>protein synthesis Aureolib</aureodatabase> | ||
</protect> | </protect> | ||
<protect> | <protect> | ||
== | ==Protein stability== | ||
* <aureodatabase>protein half-life</aureodatabase> | *<aureodatabase>protein half-life</aureodatabase> | ||
</protect> | </protect> | ||
Latest revision as of 05:28, 11 March 2016
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1514 [new locus tag: SACOL_RS07710 ]
- pan locus tag?: SAUPAN003935000
- symbol: gpsA
- pan gene symbol?: gpsA
- synonym:
- product: NAD(P)H-dependent glycerol-3-phosphate dehydrogenase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1514 [new locus tag: SACOL_RS07710 ]
- symbol: gpsA
- product: NAD(P)H-dependent glycerol-3-phosphate dehydrogenase
- replicon: chromosome
- strand: -
- coordinates: 1552712..1553710
- length: 999
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3237596 NCBI
- RefSeq: YP_186358 NCBI
- BioCyc: see SACOL_RS07710
- MicrobesOnline: 912966 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961ATGACTAAAATTACCGTTTTTGGTATGGGAAGTTTTGGGACAGCCCTTGCCAATGTTCTT
GCAGAAAATGGACATGATGTTTTGATGTGGGGTAAAAATCAAGATGCTGTTGATGAATTA
AATACATGTCATACAAATAAAAAGTATTTAAAATACGCGAAATTAGATGTTAACATCATC
GCTACTTCAGATATGACCAAAGCAATTCAATTTGCAGATATTTACTTAATGGCTTTACCT
ACTAAAGCAATGCGAGAAGTTGCTTCTCAAATTAATGATAAGCTGACCTCTAAAAAGACT
TTTATACATGTTGCTAAAGGTATTGAAAATGGGACGTTTAAACGTGTGTCAGAAATGATT
GAAGATTCTATTTCACCTGAATATAATGCAGGTATTGGCGTGTTGTCAGGGCCAAGTCAT
GCGGAAGAAGTTGTAGTCAAGCAACCAACTACAGTTGCTGCTTCATCAAAAGATAAAAGT
GTAAGTAAATTAACGCAAGATTTATTTATGAATGATTATTTGCGTGTGTACACGAATGAT
GACTTGATTGGTGTTGAACTTGGTGGTGCATTGAAAAATATCATCGCAGTAGCAAGTGGT
ATCGTAGCTGGAATTGGCTACGGTGATAATGCAAAAGCTGCATTAATGACTCGTGGCTTA
GCGGAAATTAGTAGATTAGGTGAAAAGTTAGGTGCCGATCCTATGACATTTCTAGGTTTA
GGTGGTATCGGTGACTTAATCGTTACTTGCACATCAACACATTCTCGGAATTTCACATTA
GGATATAAACTTGGACAAGGTGAATCAATGGATCAAGCATTATCTGAAATGAATATGGTT
GTTGAAGGTATTTATACAACTAAATCAGTTTATCATTTAGCTAAAGAAAAAAATGTGGAT
ATGCCAATTACAAATGCATTATATAGAGTATTATTTGAAAATATCTCAGTAAAAGAATGC
GTAAAAGATTTAATGGAGCGCGATAAAAAATCTGAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
999
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1514 [new locus tag: SACOL_RS07710 ]
- symbol: GpsA
- description: NAD(P)H-dependent glycerol-3-phosphate dehydrogenase
- length: 332
- theoretical pI: 6.26285
- theoretical MW: 36071.3
- GRAVY: -0.0858434
⊟Function[edit | edit source]
- reaction: EC 1.1.1.8? ExPASyGlycerol-3-phosphate dehydrogenase (NAD+) sn-glycerol 3-phosphate + NAD+ = glycerone phosphate + NADHEC 1.1.1.94? ExPASyGlycerol-3-phosphate dehydrogenase (NAD(P)+) sn-glycerol 3-phosphate + NAD(P)+ = glycerone phosphate + NAD(P)H
- TIGRFAM: glycerol-3-phosphate dehydrogenase (NAD(+)) (TIGR03376; EC 1.1.1.8; HMM-score: 236.2)and 8 morenucleotide sugar dehydrogenase (TIGR03026; HMM-score: 40)Biosynthesis of cofactors, prosthetic groups, and carriers Pantothenate and coenzyme A 2-dehydropantoate 2-reductase (TIGR00745; EC 1.1.1.-; HMM-score: 31.2)Energy metabolism Electron transport NADPH-dependent F420 reductase (TIGR01915; HMM-score: 24.5)Amino acid biosynthesis Glutamate family pyrroline-5-carboxylate reductase (TIGR00112; EC 1.5.1.2; HMM-score: 22.4)Energy metabolism Pentose phosphate pathway 6-phosphogluconate dehydrogenase (decarboxylating) (TIGR00872; EC 1.1.1.44; HMM-score: 22)3-hydroxyacyl-CoA dehydrogenase PaaC (TIGR02279; EC 1.1.1.-; HMM-score: 15.5)Energy metabolism Pentose phosphate pathway 6-phosphogluconate dehydrogenase (decarboxylating) (TIGR00873; EC 1.1.1.44; HMM-score: 13.6)Amino acid biosynthesis Pyruvate family ketol-acid reductoisomerase (TIGR00465; EC 1.1.1.86; HMM-score: 13.3)
- TheSEED :
- Glycerol-3-phosphate dehydrogenase [NAD(P)+] (EC 1.1.1.94)
Carbohydrates Sugar alcohols Glycerol and Glycerol-3-phosphate Uptake and Utilization Glycerol-3-phosphate dehydrogenase [NAD(P)+] (EC 1.1.1.94)and 2 more - PFAM: 6PGD_C (CL0106) NAD_Gly3P_dh_C; NAD-dependent glycerol-3-phosphate dehydrogenase C-terminus (PF07479; HMM-score: 192.5)NADP_Rossmann (CL0063) NAD_Gly3P_dh_N; NAD-dependent glycerol-3-phosphate dehydrogenase N-terminus (PF01210; HMM-score: 175)and 12 moreF420_oxidored; NADP oxidoreductase coenzyme F420-dependent (PF03807; HMM-score: 42.9)UDPG_MGDP_dh_N; UDP-glucose/GDP-mannose dehydrogenase family, NAD binding domain (PF03721; HMM-score: 35.7)ApbA; Ketopantoate reductase PanE/ApbA (PF02558; HMM-score: 33.4)NAD_binding_2; NAD binding domain of 6-phosphogluconate dehydrogenase (PF03446; HMM-score: 29.1)IlvN; Acetohydroxy acid isomeroreductase, NADPH-binding domain (PF07991; HMM-score: 23.4)3HCDH_N; 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain (PF02737; HMM-score: 23.1)TrkA_N; TrkA-N domain (PF02254; HMM-score: 17.9)2-Hacid_dh_C; D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain (PF02826; HMM-score: 15.4)6PGD_C (CL0106) ApbA_C; Ketopantoate reductase PanE/ApbA C terminal (PF08546; HMM-score: 14.9)Nucleot_cyclase (CL0276) GCH_III; GTP cyclohydrolase III (PF05165; HMM-score: 14.5)NADP_Rossmann (CL0063) Shikimate_DH; Shikimate / quinate 5-dehydrogenase (PF01488; HMM-score: 13.8)PDH; Prephenate dehydrogenase (PF02153; HMM-score: 12.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- LocateP: N-terminally anchored (with CS)
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: 0.5
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: NGHDVLMW
- SignalP: Signal peptide SP(Sec/SPI) length 21 aa
- SP(Sec/SPI): 0.614414
- TAT(Tat/SPI): 0.005277
- LIPO(Sec/SPII): 0.014003
- Cleavage Site: CS pos: 21-22. VLA-EN. Pr: 0.5386
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTKITVFGMGSFGTALANVLAENGHDVLMWGKNQDAVDELNTCHTNKKYLKYAKLDVNIIATSDMTKAIQFADIYLMALPTKAMREVASQINDKLTSKKTFIHVAKGIENGTFKRVSEMIEDSISPEYNAGIGVLSGPSHAEEVVVKQPTTVAASSKDKSVSKLTQDLFMNDYLRVYTNDDLIGVELGGALKNIIAVASGIVAGIGYGDNAKAALMTRGLAEISRLGEKLGADPMTFLGLGGIGDLIVTCTSTHSRNFTLGYKLGQGESMDQALSEMNMVVEGIYTTKSVYHLAKEKNVDMPITNALYRVLFENISVKECVKDLMERDKKSE
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Signal peptide containing [1] [2] [3] [4]
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
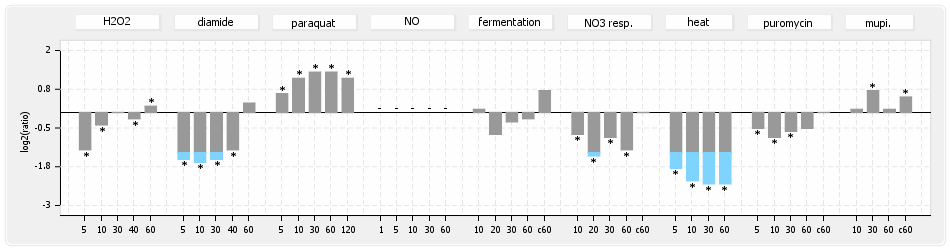
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 59.44 h [5]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You are kindly invited to share additional interesting facts.
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)